Research synopsis
The Golgi lies at the heart of the secretory pathway, receiving the entire output of newly-synthesized cargo proteins from the endoplasmic reticulum, modifying any bound oligosaccharides, and then sorting them to their final destinations. Often comprising a stack of closely-apposed and flattened cisternae, the Golgi presents a complex architecture that needs to be duplicated and partitioned every cell cycle.
The primary aim of our research work has been to understand how the cell creates another copy of the Golgi during the cell cycle and partitions them equally between the two daughter cells, thereby ensuring that this organelle is propagated through successive generations.
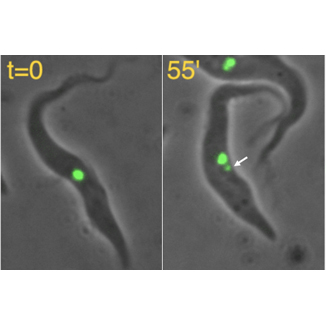
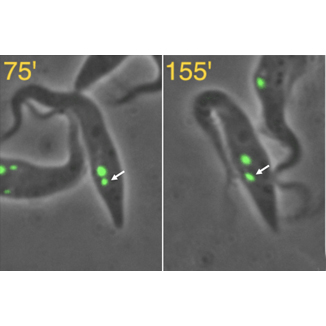
To simplify the study of this process we have focused on the protozoan parasite, Trypanosoma brucei, which has a single Golgi that undergoes duplication during the cell cycle (see Figure). Using video fluorescence microscopy and biochemical techniques, we have been able to identify novel structures that help determine the source, size, position and inheritance of the new Golgi.
Selected publications
Yavuz S & Warren G (2017). A role for Sar1 and ARF1 GTPases during Golgi Biogenesis in the protozoan parasite Trypanosoma brucei. Mol. Biol. Cell, 28, 1782-1791.
Demmel L, et al (2016). The endocytic activity of the flagellar pocket in Trypanosoma brucei is regulated by an adjacent phosphatidylinositol phosphate kinase. J Cell Sci 129.
Sealey-Cardona M, et al (2014). Sec16 determines the size and functioning of the Golgi in the protist parasite, Trypanosoma brucei. Traffic 15, 613-29.
Morriswood B, et al (2013). Novel bilobe components in Trypanosoma brucei identified using proximity-dependent biotinylation. Eukaryot. Cell 12, 356-67.
de Graffenried CL, et al (2008). Polo-like kinase is required for Golgi and bilobe biogenesis in Trypanosoma brucei. J Cell Biol. 181, 431-8.
He C, et al (2005). Golgi duplication in Trypanosoma brucei requires centrin2. Science 310, 1196-1198.
Research themes
Membrane trafficking
Biogenesis of the Golgi
 Close
Close


