Research
Noncovalent Interactions of π Systems with Sulfur: The Atomic Chameleon of Molecular Recognition
The relative strength of noncovalent interactions between a thioether sulfur atom and various π systems in designed top pan molecular balances was determined by NMR spectroscopy. Compared to its oxygen counterpart, the sulfur atom displays a remarkable ability to interact with almost equal facility over the entire range of π systems studied, with the simple alkene emerging as the most powerful partner. With the exception of the O⋅⋅⋅heteroarene interaction, all noncovalent interactions of sulfur with π systems are favoured over oxygen. For full details, see the Angewandte report of November 2017.
Tin chemical shift anisotropy in tin dioxide: On ambiguity of CSA asymmetry derived from MAS spectra
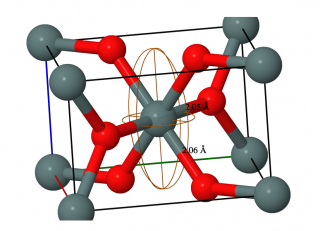
Two different axial symmetries of the 119Sn chemical shift anisotropy (CSA) in tin dioxide with the asymmetry parameter (η) of 0 and 0.27 were reported previously based on the analysis of MAS NMR spectra. By analyzing the static powder pattern, we have shown that the 119Sn CSA is axially symmetric. A nearly axial symmetry and the principal axis system of the 119Sn chemical shift tensor in SnO2 were deduced from periodic scalar-relativistic density functional theory (DFT) calculations of NMR parameters. The implications of fast small-angle motions on CSA parameters were also considered, which could potentially lead to a CSA symmetry in disagreement with a crystal symmetry. Our analysis of experimental spectra using spectral simulations and iterative fittings showed that MAS spectra recorded at relatively high frequencies do not show sufficiently distinct features in order to distinguish CSAs with η ≈ 0 and η ≈ 0.4. The example of SnO2 shows that both the MAS lineshape and spinning sideband analyses may overestimate the η value by as much as ∼0.3 and ∼0.4, respectively. The results confirm that a static powder pattern must be analysed in order to improve the accuracy of the CSA asymmetry measurements. The measurements on SnO2 nanoparticles showed that the asymmetry parameter of the 119Sn CSA increases for nm-sized particles with a larger surface area compared to μm-sized particles. The increase of the η value for tin atoms near the surface in SnO2 was also confirmed by DFT calculations. For full details, see Solid State Nuclear Magnetic Resonance of February 2018.
The structure of tagetitoxin
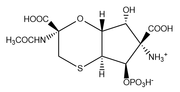
Based on detailed analysis of newly acquired NMR data, mainly long-range JCH couplings, it has been shown that the previously revised structure of tagetitoxin (bacterial phytotoxin) is incorrect. A new structure of tagetitoxin is proposed which is consistent with the NMR and MS data.
Lone pair...(no-π!)-heteroarene noncovalent interactions: the Janus faced hydroxyl group
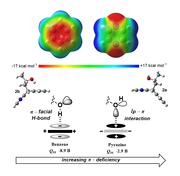
A study by NMR using designed top pan molecular balances demonstrates that the noncovalent interaction of a hydroxyl group with π-deficient pyrazine and quinoxaline units involves a lone pair...heteroarene interaction which is much stronger and essentially solvent independent when measured relative to the classical π-facial hydrogen bond to a benzene ring. Alkyl fluorides also prefer the heteroarene rings over the benzene ring. The attractive interaction between a quinoxaline and a terminal alkyne is also stronger than the intramolecular hydrogen bond to an arene. For full details, see the Angewandte report of May 2015.
Surfing π-clouds for non-covalent interactions: arenes versus alkenes
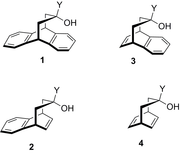
A comparative study by NMR spectroscopy using molecular balances indicates that non-covalent functional group interactions with an arene dominate over those with an alkene and a π-facial intramolecular hydrogen bond from a hydroxyl group to an arene is favoured by ~1.2 kJ mol-1. The strongest interaction observed in the Angewandte report of January 2015 is with the cyano group. Analysis of the series of CH2CH3, CH=CH2, C≡CH and C≡N groups shows a correlation between conformational free energy differences and the calculated charge of the Cα atom of these substituents, indicative of the electrostatic nature of their π-interactions. Changes in the free energy differences of conformers show a linear dependence on the solvent hydrogen bond acceptor parameter β.
Force field optimisations for biomolecular MD simulations
We have developed a new approach for force field optimisations aimed at improving the accuracy of dynamics characteristics and motionally averaged structural properties predicted by biomolecular MD simulations (Proteins 2014). As the source of experimental data for dynamics fittings, we have used 13C NMR spin-lattice relaxation times (T1) of backbone and sidechain carbons, which allow to determine correlation times of both overall molecular and intramolecular motions. For structural fittings, we used motionally averaged experimental values of NMR J couplings. The proline residue and its derivative 4-hydroxyproline with relatively simple cyclic structure and sidechain dynamics were chosen for the assessment of the new approach. Initially, grid search and simplexed MD simulations identified a large number of parameter sets which fit equally well experimental J couplings. Using the Arrhenius-type relationship between the force constant and the correlation time, the available MD data for a series of parameter sets were analyzed to predict the value of the force constant that best reproduces experimental timescale of the sidechain dynamics.
Verification of the new force-fields AMBER99SB-ILDNP and AMBER99SB*-ILDNP against NMR J couplings and correlation times showed consistent and significant improvements compared to the original force field in reproducing both structural and dynamics properties. The results suggest that matching experimental timescales of motions together with motionally averaged characteristics is the valid approach for force field parameter optimisation. Such a comprehensive approach is not restricted to cyclic residues and can be extended to other amino acid residues, as well as to the protein backbone.
The new force fields AMBER99SB-ILDNP and AMBER99SB*-ILDNP are available for downloading. These are in the format required by the Gromacs MD package.
NMR/MD/QM approach for 3D structure predictions in solutions
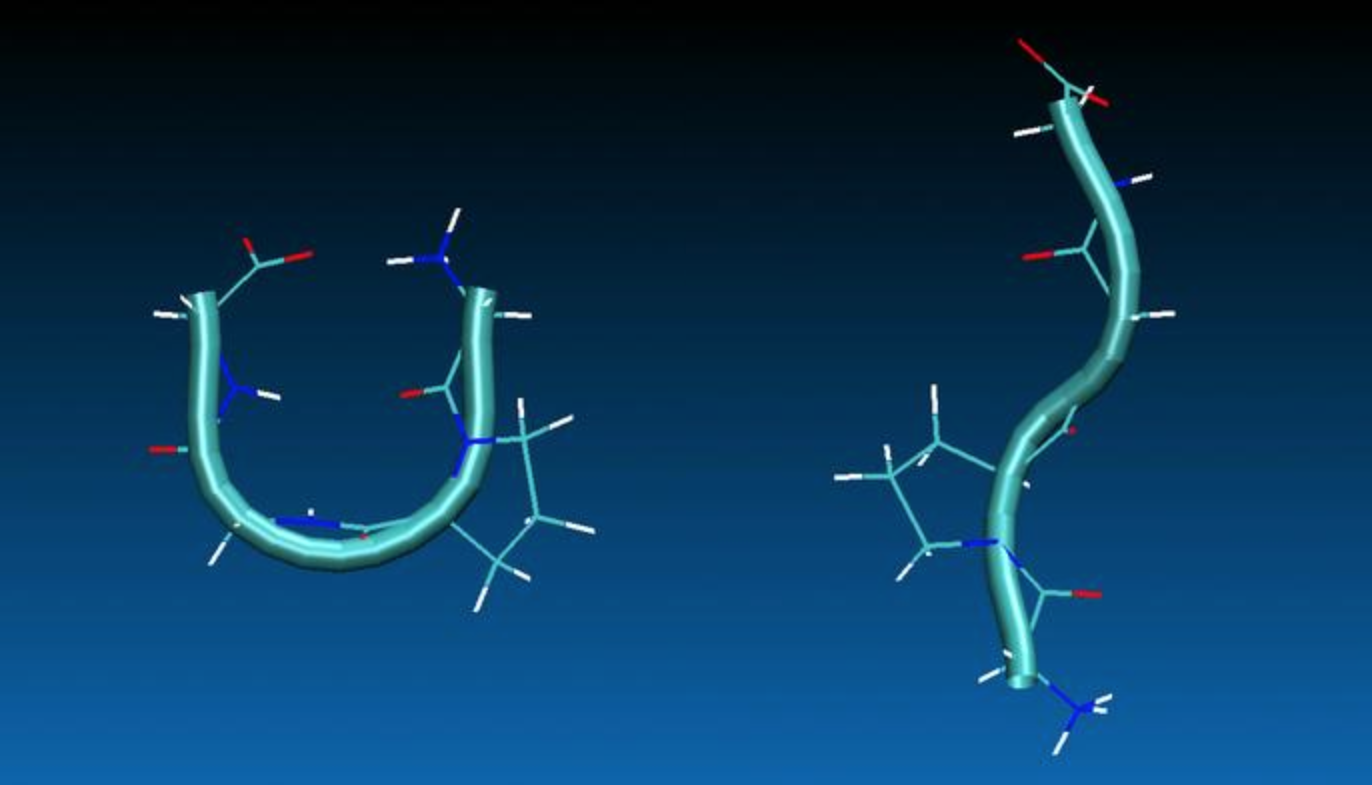
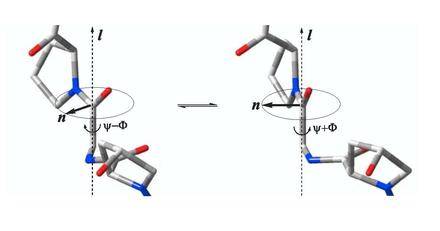
Our recent research has focused on combined application of experimental NMR techniques together with molecular dynamics (MD) simulations and quantum-mechanical (QM) calculations for structure and dynamics elucidations of small molecules in solutions. Potentially, the combined method is capable of selecting the most preferred conformations of flexible open chain molecules out of possible millions of molecular geometries. At the output, the method provides full geometric characteristics of the preferred conformations, including 3D coordinates of atoms, as in X-ray crystallography. In addition, relative populations of the major conformers are also estimated. See our paper (ChemCom 2012), in which the NMR/MD/QM technique was first introduced and then applied to tetrapeptides Gly-Pro-Gly-Gly and Val-Ala-Pro-Gly. Two main conformations of Gly-Pro-Gly-Gly were identified using NMR/MD/QM analysis, the structures of which are illustrated below.
A similar approach was also employed by us for the determination of diastereomer configurations (J Org Chem 2012).
Further verifications and applications to cyclic organic compounds are presented in J Phys Chem A 2012. Using molecules with known solid-state structures extensive analysis of various QM methods and basis sets was carried out in order to identify suitable protocols for accurate predictions of molecular geometries in solutions, as well as NMR chemical shifts and J-couplings. As the NMR/MD/QM approach relies on the final QM geometry optimization, comparisons of geometric characteristics predicted by different QM methods and those from X-ray and neutron diffraction measurements were undertaken using rigid and flexible cyclic systems. The joint analysis showed that intermolecular noncovalent interactions present in the solid state alter molecular geometries significantly compared to the geometries of isolated molecules from QM calculations.
Solid-state 2H NMR
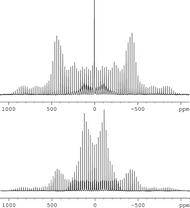
We have explored high-resolution solid-state 2H NMR and computational techniques in studies of the α and γ polymorphs of fully deuterated glycine-d5 (J Phys Chem A 2011). One of the primary objectives of this study was to prepare grounds for natural-abundance solid-state 2H NMR measurements (Chem Phys Lett 1994). In spite of the low natural abundance of the 2H isotope, the increasing availability of high-field NMR instruments creates the prospect of carrying out natural-abundance solid-state 2H NMR measurements on a routine basis, particularly for detailed characterization of structure and dynamics in solids. Shown on the right are 2H MAS NMR spectra of the α (bottom) and γ (top) polymorphs of glycine-d5 recorded on the UK 850 MHz solid-state NMR Facility. Based on the detailed analysis of these spectra using numerical methods, it was possible to characterize structural and dynamics aspects of glycine polymorphs, including differences in intra- and intramolecular interactions in two polymorphs.
Molecular dynamics in organic solids and collagen
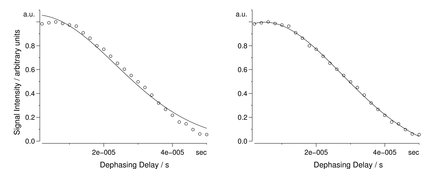
Molecular dynamics characterisations in solids can be carried out selectively using dipolar-dephasing experiments. We have shown that the introduction of a sum of Lorentzian and Gaussian functions greatly improves fittings of the "intensity vs. time" data for protonated carbons in dipolar-dephasing experiments (J Phys Chem A 2013). The Lorentzian term accounts for remote intra- and intermolecular 1H-13C dipole-dipole interactions which vary from one molecule to another or for different carbons within the same molecule. By separating contributions from weak remote interactions more accurate Gaussian decay constants, Tdd, can be extracted for directly bonded 1H-13C pairs. Reorientations of the 1H-13C bonds lead to the increase of Tdd and by measuring dipolar-dephasing constants, an insight can be gained into dynamics in solids. We have demonstrated advantages of the method using comparative dynamics studies in amino acids and peptides, including their polymorphs, as well as adamantane and hexamethylenetetramine. It was possible to distinguish subtle differences in dynamics of different carbon sites within a molecule in polymorphs and in L- and DL-forms of amino acids. The presence of overall molecular motions was found to lead to particularly large differences in dipolar-dephasing experiments. The differences in dynamics can be attributed to differences in non-covalent interactions. In the case of hexamethylenetetramine, for example, the presence of C-H…N interactions leads to nearly rigid molecules. Overall, the method allows to gain insight into the role of non-covalent interactions in solids and their influence on the molecular dynamics.
Using solid-state NMR we have shown that the amplitude of collagen backbone and sidechain motions increases significantly on increasing the water content (Biopolymers 2014). This conclusion is supported by the changes observed in three different NMR observables: (i) the linewidth dependence on the 1H decoupling frequency; (ii) 13C chemical shift anisotropy changes for the peptide carbonyl groups, and (iii) dephasing rates of 1H-13C dipolar couplings. In particular, a nearly three-fold increase in motional amplitudes of the backbone librations about C-Cα or N-Cα bonds was found on increasing the added water content up to 47 wt%D2O. Based on the frequencies of NMR observables involved, the timescale of the protein motions dependent on the added water content was estimated to be of the order of microseconds. Our wideline 1H NMR measurements revealed that the timescale of the microsecond motions in proteins reduces significantly on increasing the added water content, i.e., an approximately 15-fold increase in protein motional frequencies is observed on increasing the added water content to 45 wt%D2O. The observed changes in collagen dynamics are attributed to the increase in water translational diffusion on increasing the amount of added water, which leads to more frequent "bound water"/"free water" exchange on the protein surface, accompanied by the breakage and formation of new hydrogen bonds with polar functionalities of protein.
Links to other publications can be found here.
 Close
Close

