The cerebral cortex is essential for our most advanced mental abilities, such as language, complex motor skills and abstract thinking. While the cortex is larger in humans than other animals, its basic "circuit diagram" appears largely the same, suggesting that evolution has reprogrammed a general-purpose cortical circuit to perform new and more complex tasks. Understanding this circuit is therefore of paramount importance to neuroscience.
Cortical circuits, however, are built from a great diversity of cell types, and understanding cortical computation will require identifying the role each type plays in circuit function and in behavioral tasks such as sensory perception. Techniques such as two-photon microscopy allow one to record the activity of thousands of cells simultaneously in behaving subjects. Moreover, using laser light to induce individual neurons to fire, we can learn how each cell in the circuit influences the activity of the others.
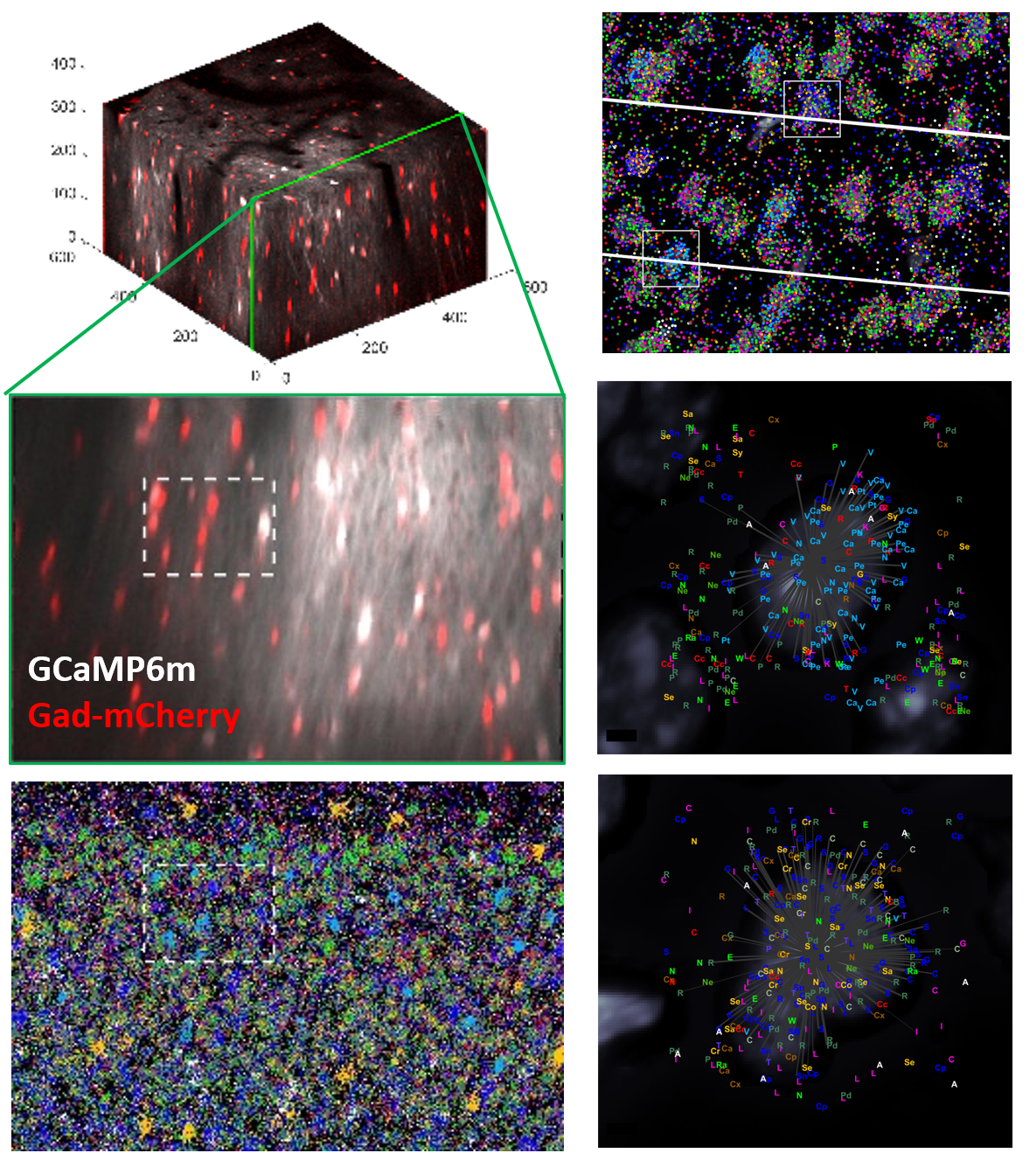
However, it has not yet been possible to identify the types of more than a few of these recorded cells. Transcriptomics offers a solution to this problem. Single-cell RNA sequencing allows one to rigorously define cell types by multivariate analysis of gene expression levels in thousands of individual cells, and to identify marker genes for each type. In situ RNA sequencing allows these markers to be simultaneously localized in cell populations whose activity was previously recorded by two-photon calcium imaging. Together, these methods therefore allow simultaneous recording of thousands of cells, with post-hoc identification of each cell's fine transcriptomic type. Our first application of this method (published here) found that the main functional difference between V1 interneurons is not their sensory tuning, but their modulation by brain state, and found that despite the tremendous complexity of these cell types, a single transcriptomic axis can account for most of their differences in state modulation. Current work is applying this method to understand how fine cortical cell types operate during active behaviors, and how they are modulated by learning. Additionally we are using the methods to study models of disease conditions such as Alzheimer's, in collaboration with the UK Demential Research Institute.
The development of this method was funded by a strategic award from the Wellcome Trust from 2015 to 2020, to a multidisciplinary team involving the research groups of Kenneth Harris, Matteo Carandini, Michael Hausser, Bill Richardson, Nicoletta Kessaris, Peter Somogyi, Mats Nilsson, Jens Hjerling-Leffler and Sten Linnarsson, at UCL, Oxford University, and Karolinska Institutet. Some other publications from this collaboration are here and here.
To see data from single-cell RNA sequencing, see here. To see data from in situ sequencing of hippocampus in an online viewer, see here.

 Close
Close