Research Themes
Transcription and RNA Biology
Regulating the genetic flow is the key to survival, adaptation and the body’s response to disease. Genetic information is stored and regulated across different layers of DNA and RNA organisation and function. We ask how RNA is made and processed and how the regulatory layers are connected and respond to changes in the cellular and extracellular environment. Using model systems from the different domains of life, we study the pathways regulating the synthesis, metabolism and translation of messenger RNA molecules in space and time. The knowledge we have gained defines mechanisms underlying pathologies ranging from viral and parasitic infection to cancer and neurodegeneration.
Metabolism and Disease
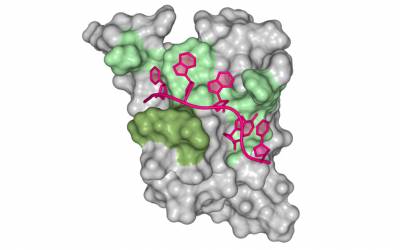
The chemistry of life builds simple molecular events into pathways of cellular and molecular complexity. We study metabolic mechanisms underlying the development, growth and function of various cell types. We investigate metal homeostasis, stress response and the antioxidant response in both health and diseases including cancer, immune disorders and neurodegeneration. We combine modern biochemistry with cellular, structural and molecular methodologies to advance our research programs. Importantly, our approach requires the development of novel optical techniques to investigate the chemical events of metabolism in vivo. Ultimately, our goal is to modulate metabolic and redox homeostasis to maintain good health, and treat diseases associated with their imbalance.
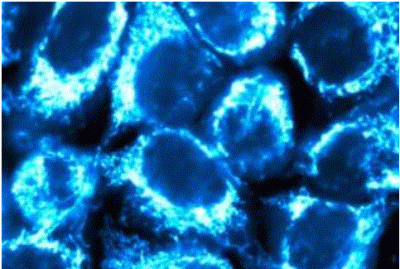
Membranes, Signalling and Trafficking
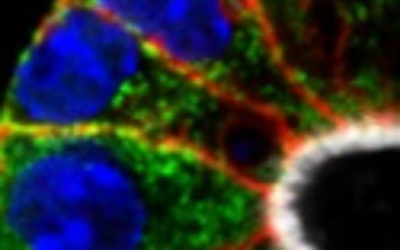
Eukaryotic cells are divided into many different organelles by membranes, creating distinct compartments, each performing a unique function. Each organelle requires specific components to perform its function. We study how proteins are targeted to the correct place within the cell, and how these processes are regulated. We determine how the cell responds to various extracellular signals, and how pathogens exploit cellular systems. We also investigate what happens when membrane trafficking and signalling machineries go wrong, leading, for example, to neurodegenerative diseases, diabetes or cancer.
Protein folding in health and disease
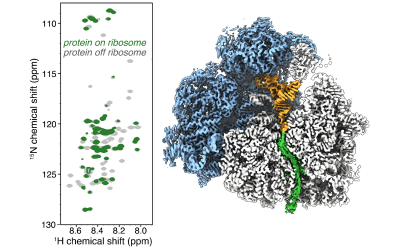
Proteins are the cell’s workhorses, and their unique 3D structures and motions underlay their cellular function(s). We study how proteins fold and acquire their dynamic structures in cells, how their structures are maintained, and the interactions that they make with proteins, nucleic acids and co-factors for biological activity within cells. We also examine how and when protein folding fails, to understand the pathogenesis of an increasingly wide range of human diseases including neurodegeneration, cancer and respiratory diseases.
Microbes and Environment
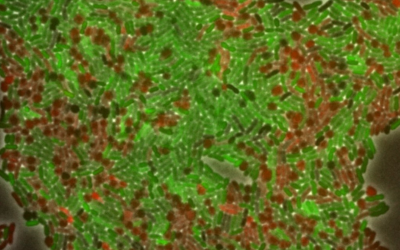
Pathogenic microbes are the main cause of human disease and the source of recurring global heath emergencies. Equally important, microbial populations provide the basis for life, and have great potential in the production of biomass, metabolites and proteins. We study how microbes interface with each other and the environment and how they evolve these capabilities, but also how we can re-engineer them to take advantage of their potential. We combine structural methods and biophysics, with advanced microscopy, omics and genetic engineering and understand the microbes’ gene regulation, metabolism and population dynamics in the host.
Computational Biology and AI
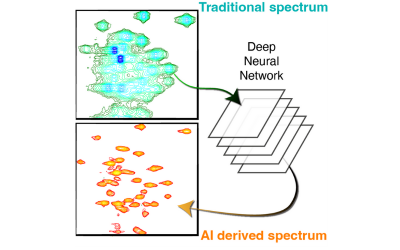
Computational biology lies at the core of modern biological research. In SMB, groups with wide-ranging interests from molecular dynamics through molecular orbital methods to artificial intelligence and deep learning, work on a broad range of subjects, from protein classification and function prediction, cell and molecular dynamics, to antibody informatics. The SMB community develops AI tools to track cells and particles within cells, and to increase the scope of biological spectroscopic methods, among many other areas. The methods, tools and databases we are developing cover a wide range of applications in the broader area of computational biology and provide new discovery pathways in biological research.
 Close
Close