CLRN2 confirmed as new deafness gene
1 February 2021
New study to impact future diagnosis and treatment for deaf patients.
A new article "A biallelic variant in CLRN2 causes non-syndromic hearing loss in humans", published online last week in Human Genetics (Springer Nature) identifies CLRN2 as a new autosomal recessive deafness gene in humans and zebrafish and describes a new deafness-causing allele in mice.
Animal studies uncovered that clarin 2 is essential for normal organization and maintenance of mechanosensitive hair bundles. This study shows the first family to segregate a likely pathogenic variant in CLRN2. Affected individuals in the family report moderate-to-profound sensorineural hearing loss.
The identification of additional families will be crucial to refine and dissect clinical aspects of hearing loss due to variants in CLRN2.
Lead author Barbara Vona from the Department of Otolaryngology, Head and Neck Surgery at the University of Tübingen, Germany, said:
“Since the completion of the Human Genome Project, researchers have the enormous task to characterize essential genes in the human genome that are crucial for normal hearing. One such way to identify these genes is to analyze seqencing data of large families with hearing loss and pinpoint strong variants that may have a damaging consequence the coding protein. The strong variant that we found in CLRN2 set the stage for international collaborative efforts and led to the discovery of a new hearing loss gene.
UCL Ear Institute's Dr Mike Bowl, co-senior author of the study, said:
“Hearing loss is an extremely common condition, and while hearing aids and cochlear implants benefit some people, they do not restore hearing or work for everyone. In the future, it may be possible to use gene-based technologies as a therapy. However, this will only be possible if we have a full understanding of the molecular processes that underlie hearing. Human/mouse cross-validation studies, as undertaken in this international collaborative project, will continue to elaborate upon the genetic landscape of hearing and provide insight to the molecular mechanisms and pathobiology of deafness.
Image
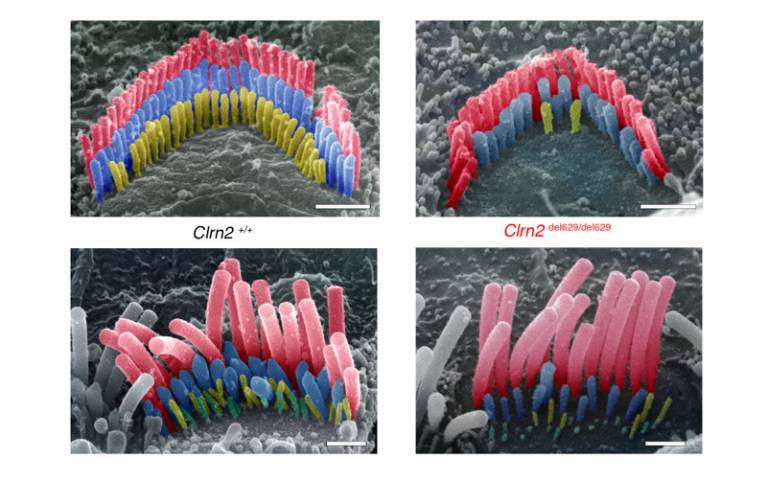
- Pseudo-colored scanning electron micrographs from the Clrn2+/+ (normal hearing) and Clrn2del629/del629 (hearing loss) mouse cochlea illustrate the three rows, tallest (red), middle (blue) and short (yellow), of P28 (± 1 day) stereocilia in inner and outer hair cell bundles. Lack of clarin 2 causes complete or partial regression of the short row stereocilia. (Scale bar = 1 μm)
Links
- Dr Mike Bowl
- Bowl Lab
- Link to full paper on Springer Nature's Human Genetics
- Dr Barbara Vona profile on researchgate
- Dr Barbara Vona Research Group
 Close
Close

