Surface Plasmon Resonance (SPR) & Dual Polarisation Interferometry (DPI)
BiacoreX100 surface plasmon resonance instrument
Surface plasmon resonance (SPR) studies of protein in the solution phase flowing over and binding to an immobilised protein surface enables the protein association to be characterized. As the analyte is flowed over the immobilized partner, the response increases, giving the “on-rate”, and when buffer is then flowed over the partner, the response decreases, giving the “off-rate”. The extent to which different molecules interact with a single partner immobilized on a sensor surface reveals the specificity of an interaction. Apart from the need to immobilise one partner on a chip surface, the method is artefact-free and does not require the use of any labelling.

The ratio of the on-rate and off-rate gives the dissociation constant KD value, which measures the strength of the binding. If the on-rate and off-rate are rapid, only the overall response is measured in what are termed equilibrium measurements. Here interaction stoichiometry is measured from comparing the increase in response units to the amount of response units immobilised on the sensor surface.
Farfield dual polarisation interferometer
Macromolecules can change shape when immobilised on a sensor surface and then subsequently treated with different buffers, small ligands or other macromolecules. Unlike surface plasmon resonance, these shape changes are measurable using a dual polarisation interferometer. This approach can often provide vital clues on bimolecular mechanisms, especially when considering interactions involving proteins immobilised on cell surfaces. Like surface plasmon resonance, information on dissociation constants is also obtained.
Interaction between Factor H and Complement C3b
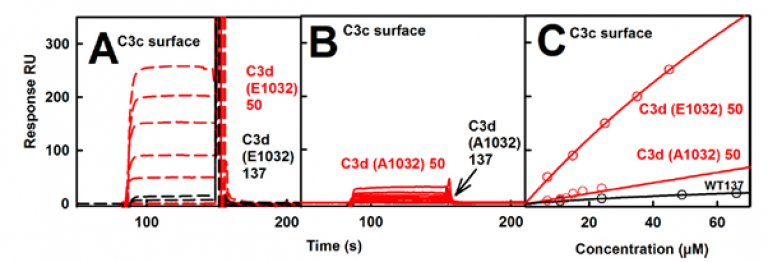
Complement C3b exists in two allotypes, namely C3S and C3F, containing Arg102 and Gly102 respectively. C3S is wild-type while C3F is disease-linked. Solution scattering showed that C3b has a compact structure in low salt and a more open structure in normal salt. Modelling of these two structures showed that the TED and MG1 domains were connected through an Arg102-Glu1032 salt bridge, thus explain the two different structures. Surface plasmon resonance was used to monitor the binding of wild-type C3d(E1032) and mutant C3d(A1032) to immobilized C3c.The wild-type C3d bound to the C3c surface (A below), while the mutant C3d did not (B below). The molecular basis for the known clinical associations of the major C3S and disease-linked C3F allotypes of C3b were experimentally explained for the first time by this work.
Publication:
Rodriguez, E., Nan, R., Li., K., Gor, J. & Perkins, S. J. (2015). A revised mechanism for the activation of complement C3 to C3b: a molecular explanation of a disease-associated polymorphism. J. Biol. Chem. 290, 2334-2350. Pubmed 25488663 doi: 10.1074/jbc.M114.605691
Self-association between Factor H at Surfaces
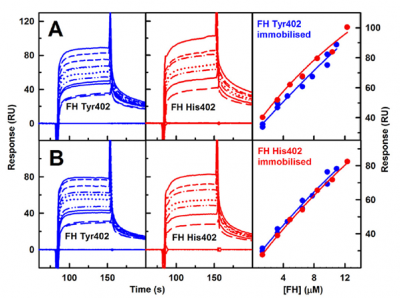
Factor H self-association is relevant for analysing its interactions with C3b, heparin and other ligands. Surface plasmon resonance experiments showed that the wild-type Tyr402 and the disease-risk His402 allotypes of Factor H self-associated with each other with similar affinities. This confirmed the results with analytical ultracentrifugation, showing that Factor H self-association has a noticeable role.
Publication:
Osborne, A. J., Nan, R., Miller, A., Bhatt, J. S., Gor, J. & Perkins, S. J. (2018) Two distinct conformations of factor H regulate discrete complement-binding functions in the fluid phase and at cell surfaces. J. Biol. Chem. 293, 17166-17187.
 Close
Close

