Functional Gene Annotation
Lead: Professor Ruth Lovering
Students: Renzhi Su, Qianhan Xu, Siyao (Nancy) Huang
Volunteer biocurators: Marios Makris, Sandra De Miranda Pinheiro, Alice Dashow
We curate the scientific literature to provide;
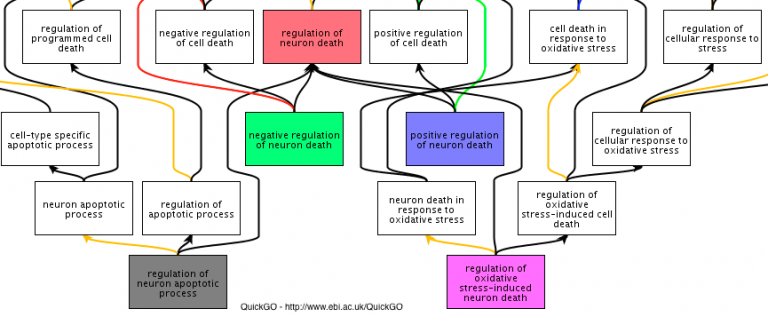
Gene Ontology (GO) annotations to the GO Consortium
Protein interaction data to the IMEx Consortium

The Functional Gene Annotation team is supported by the National Institute for Health Research University College London Hospitals Biomedical Research Centre. We support the work of the International Society for Biocuration (ISB).
 | |
 Close
Close








