The Neurological Gene Annotation project is focused on Gene Ontology (GO) annotation of proteins and microRNAs relevant to neurological processes and on contribution of new GO terms to the GO database
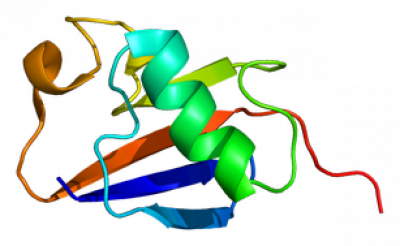

Annotations and GO terms contributed by this project to the GO Consortium database are attributed to ParkinsonsUK-UCL, ARUK-UCL or SynGO-UCL. The human LRRK2 protein record at QuickGO provides an example of a curated protein.
By using GO to curate the scientific literature we are creating a resource for the Parkinson's, Alzheimer's and neurological research communities that will enable researchers to rapidly evaluate and interpret existing data and generate hypotheses to guide future research.
Projects

We are using the GOA-UniProt curation tool to associate Gene Ontology (GO) terms with neurological-relevant proteins and microRNAs; a process known as GO annotation. This data is then incorporated into the GOA-UniProt and GO Consortium databases. All of our annotations are then propagated to freely available online knowledgebases, such as UniProt, Ensembl, RNAcentral and NCBIGene, as well as other public and commercial analysis tools. There are many ways in with the impact of our curation on data analysis and annotation resources can be demonstrated.
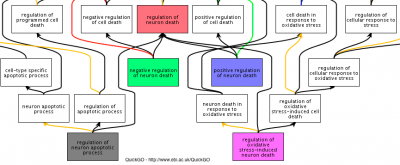
One of the approaches we take is to annotate one biological process at a time, rather than one gene product at a time. This approach has several advantages, including: broadening our annotations to include more of the proteins associated with the process, improving the biocurator's knowledge of the biology and creating more specific annotations. In turn, this leads to requests for more specific GO terms, because we are investing in improving the annotation of the domain.
What we curate
- Alzheimer's disease-relevant processes
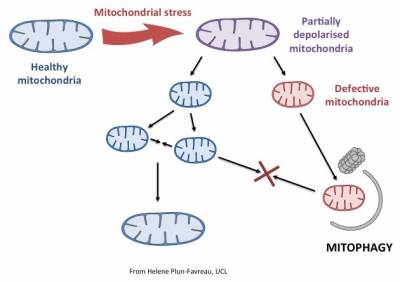
- Parkinson's disease-relevant processes
- Synapse associated proteins
- Mendelian genes associated with autism (MSc project)
To find out more about Alzheimer's or Parkinson's diseases, please see the Alzheimer's Research UK or Parkinson's UK websites.
 Close
Close










