UCL project tracking viruses through time and space wins at IET Awards
27 November 2018
A major five-year project has won the prestigious Healthcare Technologies award at this month’s Institution of Engineering and Technology (IET) Awards.
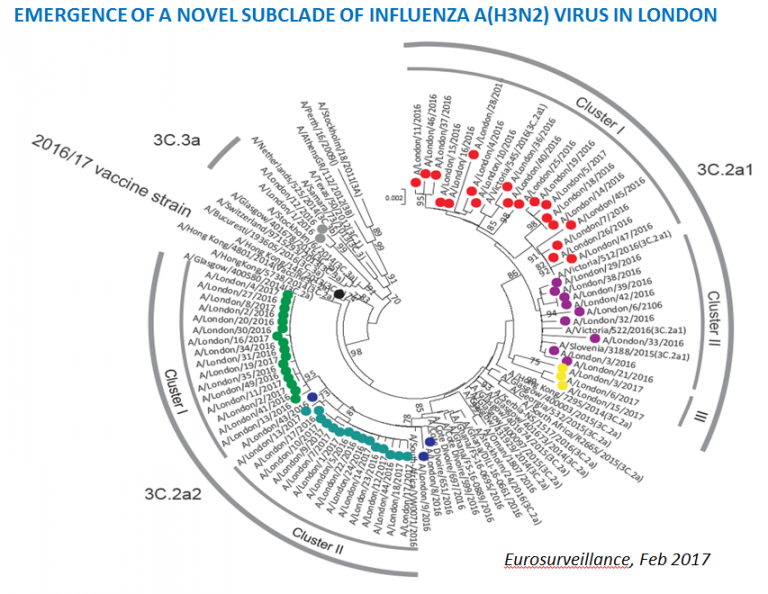
ICONIC (Infection response through virus genomics), was a flagship Wellcome Trust and National Institute for Health Research co-funded project, hosted at UCL and now translated into the University College London Hospital (UCLH) Advanced Pathogen Diagnostics Unit.
The project (2013-2018) developed the first whole genome sequencing of viral pathogens from a tool used only in basic research to one integrated into routine NHS healthcare services. This has had significant impact in stratifying patients to improve care and in controlling the spread of infectious outbreaks.
Viral infections, such as HIV, influenza (flu) and norovirus, are a serious burden to the NHS with influenza pandemics alone costing in excess of £2billion annually. Infectious outbreaks in hospitals are a particular concern due the vulnerability of patients with compromised immune functions, leaving them at greater risk of severe symptoms and death.
The ICONIC project combines the genetic data from the virus together with electronic health records to investigate and visualise these outbreaks and help bring them under control.
This pipeline was used to successfully support the control of a major hospital outbreak of influenza. The outputs of this work have been adopted into clinical practice and routine healthcare in numerous NHS settings and through various projects, supported by UCL GEO, in four different low-and-middle-income country settings.
Principle investigator Professor Andrew Hayward (Director, UCL Institute of Epidemiology and Health Care) commented, “This was an immense interdisciplinary team effort involving multiple academic and NHS partners led from the UCL Institute of Health Informatics. The work demonstrated the value of combining virus genomic data with electronic health records to investigate outbreaks and help bring them under control. We are truly delighted that this team effort has been recognised by this prestigious award.”
Clinical lead Dr Eleni Nastouli (Head of Virology, UCLH) said, “We are honoured to receive this award alongside our colleagues. As the main clinical site, UCL further developed protocols for whole genome sequencing of viruses and provided exemplars from our clinical practice of how these can be used to improve patient care and outcomes. We are now taking these ideas forward establishing the UCLH Advanced Pathogen Diagnostics Unit, a late translational research unit within the Health Services Laboratories, a UCLH partnership. As we aim to ensure patient access to novel diagnostics this award adds to our commitment and enthusiasm.”
Dr Zisis Kozlakidis who led on the implementations abroad (now Head of Laboratory Services and Biobanking, WHO/IARC) agreed, “We are proud to have addressed a significant clinical need with a team consisting of many partners across industry, academia and hospitals. There are two major breakthroughs here: firstly, creating an effective solution for the NHS and secondly, implementing and testing this experience in many locations abroad, especially in low-and-middle-income countries where the need is most acute. These projects demonstrate that effective technology transfer to these regions is not just possible, but a realistic proposition.”
The 2018 IET Awards were held on 14 November and featured over 400 top innovators from across the spectrum of engineering technology. The ICONIC project won in the Healthcare Technologies category.

The project was led from the UCL Institute of Health Informatics and was a truly interdisciplinary effort, with main clinical partners at University College London Hospital, Barts Health NHS Trust and other NHS Trusts, private partners Intel (Health Intelligence Unit) and Illumina, sequencing and bioinformatics input from the Sanger Institute and University of Edinburgh, and collaborative support from multiple other academic institutions. It was funded as a flagship Healthcare Innovation Challenge project by the UK Department of Health and the Wellcome Trust.
Related reading:
Harvala H, et al. (2017) Emergence of a novel subclade of influenza A(H3N2) virus in London, December 2016 to January 2017. Euro Surveill.;22(8):pii=30466
Houlihan CF, Frampton D, et al. (2018) Use of Whole-Genome Sequencing in the Investigation of a Nosocomial Influenza Virus Outbreak, J of Infect Dis, jiy335, https://doi.org/10.1093/infdis/jiy335
Sandmann F, et al. (2018) Estimating the true hospital burden of norovirus-associated gastroenteritis in England and its opportunity costs for non-admitted patients. Clin Infect Dis, ciy167, https://doi.org/10.1093/cid/ciy167
Smith CM, Kozlakidis Z, Frampton D, et al. (2017) Development of a novel application for visualising infectious diseases in hospital settings. The Lancet; 390(3): S84
Vandenberg O, Kozlakidis Z, et al. (2018) Control of Infectious Diseases in the Era of European Clinical Microbiology Laboratories Consolidation: New challenges and opportunities for the Patient and the Public Health surveillance. Front. Med., https://doi.org/10.3389/fmed.2018.00015
Yebra G, et al. (2018) A High HIV-1 Strain Variability in London, UK, Revealed by Full-Genome Analysis: Results from the ICONIC Project. PLoS One, 13(2):e0192081.
 Close
Close

