News about the mechanisms and evolution of transcription!
1 October 2021
Fabian Blombach and Simona Pilotto, from the RNA Polymerase Lab at the ISMB, are lead authors of two breakthrough papers in Nature Communications, studying the mechanisms and evolution of transcription.

Transcription is carried out by evolutionary conserved RNA polymerases and subject to regulation by different strategies. The control of individual genes is enabled by a plethora of DNA-binding transcription factors that respond to changes in the environment and enable the up- or downregulation gene expression and the structural basis and mechanisms of gene-specific regulation have been characterised in great detail. Much less is known about the global regulation of transcription that is enabled by RNAP-binding factors.
Two research articles from Professer Finn Werner's RNAP laboratory at the ISMB, published back-to-back in Nature Communications, explore and characterise the global control of the archaeal RNA polymerases in two different scenarios. Firstly, a multidisciplinary study combining functional genomics with in vitro transcription assays hints at a paradigm shift for the regulation of transcription in Archaea (Blombach et al., 2021), and secondly, a cryo-EM analysis elucidates, for the first time, the structural basis of RNAP inhibition by repressors involved in the archaeal virus-host arms race (Pilotto et al., 2021).
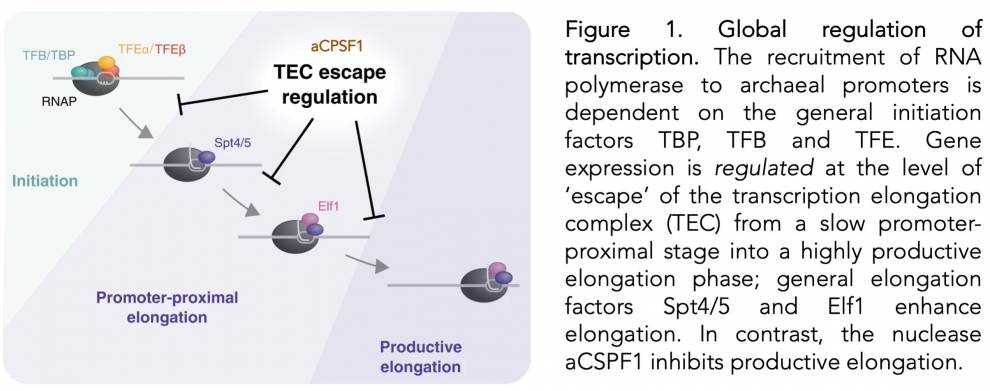
Dr Fabian Blombach, the lead author of the first study explains: ‘The current paradigm in the field dictates that transcription regulators interfere with the recruitment of RNA polymerase to the promoter at the stage of initiation. We challenged this hypothesis by comparing the profiles of RNA polymerase and transcription factors on each gene with the respective RNA level in the cell. We could show that it is not only simple ‘access’ of RNAP to the promoter that determines the expression level of a gene, but sophisticated mechanisms that occur during early transcription elongation duiring the recruitment of elongation factors Spt4/5 and Elf1 and before the RNA polymerase makes a transition into productive transcription. (Fig. 1). Quite surprisingly, we also found that RNA polymerases in the early elongation phase recruit a ribonuclease, the transcription termination factor aCPSF1. This finding suggests that premature termination emerged in parallel evolution as a regulatory strategy in the three domains of life.’
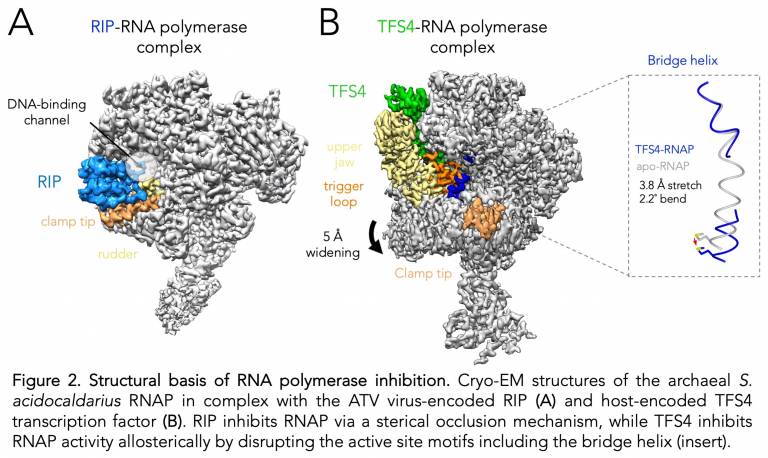
Dr Simona Pilotto, the lead author of the second study elaborates on her work: The inhibition of RNAP and the resulting attenuation of the transcriptome plays a crucial role in the interaction between viruses and their hosts during infection. My aim is to understand the structural basis of ‘switching off’ RNAP, as this is highly relevant for the design of novel antibiotics and antiviral drugs. I have solved the cryo-EM structures of the complexes formed between the Sulfolobus acidocaldarius RNAP and two distinct regulators (RIP and TFS4), and my structures provide insights into the detailed mechanisms underlying the very potent inhibition (Fig. 2). RIP is encoded by the Acidianus two-tailed virus (ATV) and sterically interferes with the interaction of template DNA and transcription factors by molecular mimicry of initiation factors. TFS4 is encoded by the Sulfolobus host and is expressed in response to infection with the Sulfolobus Turreted Icosahedral Virus (STIV). TFS4 is related to elongation factors including TFIIS and targets the RNAP NTP entry pore through which it inactivates RNAP in an allosteric fashion by inducing a widening of the DNA-binding channel disrupting two key motifs within the active site, termed 'bridge helix' and 'trigger loop'. The most intriguing conclusion of my work is that the inhibitory strategies and mechanisms reveal the underlying functional conservation of RNAPs: unrelated inhibitors have evolved to exploit factor- and nucleic acid binding sites, and conformational flexibilities that are intrinsic to all RNAPs to effectively repress its activity.
 Close
Close

