In nuclear medicine studies, the uptake, retention and washout of a tracer in a tissue region depends on physiological and biochemical parameters, such as blood flow, metabolic rate and receptor concentration. These parameters can be quantified by analysing time-activity curves obtained from dynamic PET or SPECT scans - a procedure known as kinetic modelling.
At the INM we have a number of ongoing projects that based on kinetic modelling. Some of these involve development of new methods by combining PET and MRI data (1,2 and 3 below), some involve optimization of acquisition and analysis methods for particular tracers (2 and 3).
1) Arterial Input Function estimation
A good measurement of the arterial input function (AIF) is required to estimate kinetic parameters accurately. The current gold standard method requires collection of arterial blood samples, which makes it impractical due to its invasiveness. Imaged derived input function (IDIF) is an alternative method used to extract the AIF from reconstructed PET images, but has some limitations due to the restricted spatial resolution of PET scanners.
The main goal of this research is to develop non-invasive PET input function measurement techniques utilising MRI data. A practical IDIF method for brain images has been developed which only requires segmentation of the carotid arteries from TOF MR Angiography images [1]. A novel partial volume correction (PVC) method, called Single Target Correction method has been introduced, which is tailored for IDIF extraction methods as it only requires segmentation of the carotid arteries with no need for additional parcellation of background regions. In addition, the information from partial volume corrected whole-blood TACs were combined with the simultaneous estimation approach to non-invasively analyse PET tracers where radiometabolites are present in the blood plasma. This methodology has been tested with simulated FDG data [2] and 11C-SB207145 PET tracer [3] which is used for serotonin 4 receptor (5-HT4 receptor) studies and the input function needs to be corrected for metabolites present in the plasma.
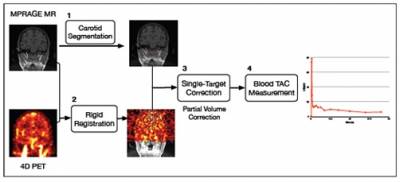
Figure 1: Brief description of IDIF extraction method, including segmentation, registration and PVC.
Publications
[1] H Sari, et al., Estimation of an image derived input function with MR-defined carotid arteries in FDG-PET human studies using a novel PVC method, 2016. J Cereb. Blood Flow Metab., in press
[2] H Sari, et al., Incorporation of MRI-AIF information for improved kinetic modelling of dynamic PET data, 2015. IEEE Transactions on Nuclear Science, 62 (3), 612-618. DOI: 10.1109/TNS.2015.2426952
[3] H Sari, et al., Blood free modelling of PET tracers with metabolites using MRI corrected whole blood and a constrained simultaneous estimation approach, PSMR Conference, Cologne, Germany, May 2016
2) ASL-incorporated kinetic modelling of PET data with reduced acquisition time
Pharmacokinetic analysis of PET data typically requires at least one hour of image acquisition, which poses a great disadvantage in clinical practice. In this work, we
propose a novel approach to perform pharmacokinetic modelling with significantly reduced PET acquisition time, by incorporating the blood flow information from simultaneously acquired arterial spin labelling (ASL) MRI. Evaluation on clinical amyloid imaging data from an Alzheimer's disease study shows the proposed approach with simplified reference tissue model can achieve amyloid burden estimation from 30-min [18F]florbetapir data and 5-min ASL MR data acquired simultaneously which is comparable with the estimation from 60-min [18F]florbetapir data.
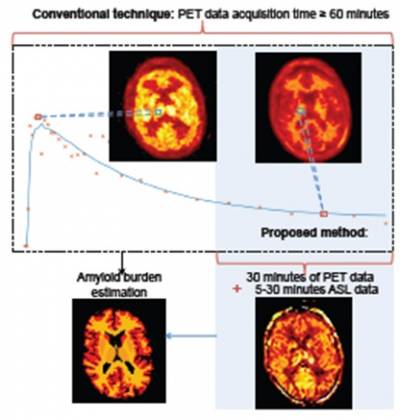
Reference: Scott C et al. (2016) MICCAI, Athens, Greece.
Figure2: Schematic of conventional dynamic PET acquisition time for pharmacokinetic modelling to estimate amyloid burden and the reduced time required for the proposed method.
3) Improved parameter-estimation with MRI-constrained PET kinetic modelling
We have investigated the potential of MRI-constrained PET kinetic modelling using simulated [18F]2-FDG data. The volume of distribution, Ve, for the extra-vascular extra-cellular space (EES) can be estimated by dynamic contrast enhanced (DCE) MRI, and then used to reduce the number of parameters to estimate in the PET model. We used a 3 tissue-compartment model with 5 rate constants (3TC5k), in order to distinguish between EES and the intra-cellular space (ICS). The results from the simulations showed reductions in bias and variance with Ve constraint. The accuracy of the parameters estimated with our new modelling approach depends on the accuracy of the assumed Ve value. In conclusion, we have shown that, by utilising information that could be obtained from DCE-MRI in the kinetic analysis of [18F]2-FDG-PET data, it is in principle possible to obtain better parameter estimates with a more complex model, which may provide additional information as compared to the standard model.
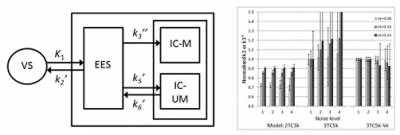
Figure 3: 3TC5k model (left), and results obtained with different models (right), showing reduced bias and variance using this model with fixed Ve.
Reference: Erlandsson K, et al., (2016) IEEE Trans. Nucl. Sci.. in press, DOI: 10.1109/TNS.2015.2507444
4) Quantification of [18F]-FMISO binding in colorectal tumours
In solid tumours, hypoxic regions can develop due to poor blood supply. Hypoxia is associated with poor treatment outcome. The PET tracer [18F]-FMISO can be used to map hypoxic tumour regions. Under hypoxic conditions, FMISO binds irreversibly to macromolecules and becomes trapped inside the cells. Due to the poor blood supply, tracer molecules typically need to diffuse across large distances before reaching the hypoxic region. This results in a low initial uptake, followed by slow accumulation of tracer. Well perfused regions, on the other hand, will show high initial uptake, followed by wash-out. As a result, the two time-activity curves may cross at some time-point. For this reason, a dynamic imaging protocol may be advantageous compared to a single late static scan, which has been used in the past. We have analysed dynamic FMISO data from patients with colorectal tumours, and compared various different data acquisition protocols and kinetic modelling approaches [1,2]. Some results are shown below.
![Parametric [18F]-FMISO images (left) and time-activity curves for hypoxic and normal VOIs (right) Parametric [18F]-FMISO images (left) and time-activity curves for hypoxic and normal VOIs (right)](https://www.ucl.ac.uk/nuclear-medicine/sites/nuclear_medicine/files/styles/medium_image/public/Kinetics4.jpg?itok=zKD9AdWC)
Figure 4: Parametric [18F]-FMISO images (left) and time-activity curves for hypoxic and normal VOIs (right).
References:
[1] Erlandsson K et al., (2016), Quantification of [18F]-FMISO binding in colorectal tumours using dynamic PET data: VOI- and voxel-based approaches, SNM annual meeting, San Diego, Ca.
[2] Erlandsson K et al., (2016), A simplified acquisition protocol for dynamic [18F]-FMISO PET studies of colorectal tumours, SNM annual meeting, San Diego, Ca.
 Close
Close

