Superbug genome report
18 July 2006
The emergence of 'superbugs' - bacteria resistant to several antibiotics - is a major problem facing healthcare providers worldwide.
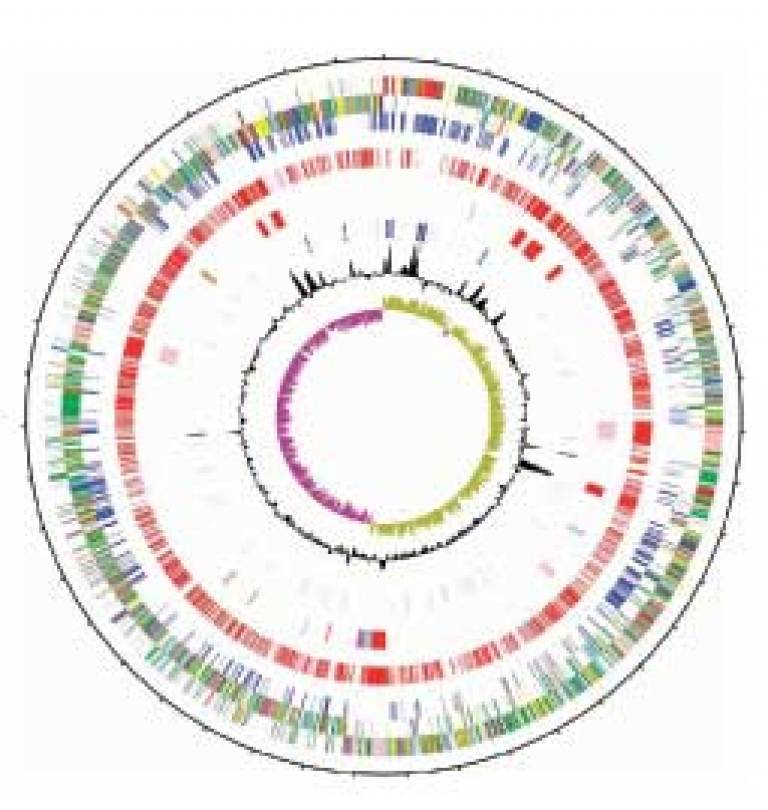 A collaborative team of geneticists have published the genome of the multi-antibiotic-resistant superbug, Clostridium difficile (C. diff) in the July 2006 issue of 'Nature Genetics'.
A collaborative team of geneticists have published the genome of the multi-antibiotic-resistant superbug, Clostridium difficile (C. diff) in the July 2006 issue of 'Nature Genetics'.
Led by the Wellcome Trust Sanger Institute, the project includes Dr Peter Mullany and Dr Adam Roberts (UCL Eastman Dental Institute Division of Microbial Diseases), and scientists from Imperial College London, the London School of Hygiene and Tropical Medicine, University of Nottingham and the Institut Pasteur in Paris.
C. diff is the leading cause of hospital-acquired infections in the developed world and caused more than 44,000 cases of disease across the UK in 2004. C. diff has become even more prevalent and is responsible for a greater mortality rate than the notorious superbug MRSA.
The research shows that half of the genes found in C. diff are absent from four of its bacterial cousins - the three species that cause botulism, gas gangrene, tetanus plus one harmless species, Clostridium acetobutylicum - and that even C. diff strains themselves are highly variable. Most important, and unlike its nearest relatives, C. diff can readily exchange genes and resistance elements. The team emphasises that this research "serves as a timely reminder on restricting the use of antibiotics given orally".
C. diff is an anaerobe - it thrives in the absence of oxygen - and can 'hibernate' in adverse conditions by forming spores. It is thought that people are most commonly infected by picking up C. diff spores. Because spores are highly resistant to most disinfection methods, C. diff is very difficult to eradicate and can easily spread among patients.
The report states that C. diff can thrive when patients are given broad-spectrum antibiotics, which wipe out protective gut bacteria, allowing C. diff to multiply and release its toxins. It causes a range of diseases from antibiotic-associated diarrhoea to a life-threatening colon disease, called pseudomembranous colitis.
C. diff is resistant to several antibiotics and is treatable with only two antibiotics, metronidazole and vancomycin. However, there are well-founded fears that it may become resistant to these.
Because C. diff is difficult to study in the lab, many of the discoveries emanating from the genome sequence are completely novel and will provide insights into the mechanisms by which the organism causes disease, help in the development of new strategies to detect and prevent infections, and may ultimately lead to new treatments.
Image: C. diff genome sequence (Nature Publishing Group)
 Close
Close

