Linguistic AmPs
"A
linguistic model for the
rational design of antimicrobial peptides" belongs loosely within a
respectable research tradition of attempting to understand complex
molecular biological systems -- in particular, genes and their products
-- in terms of language.
Genes encode information and are richly expressive, encompassing (to
leading order) the breadth and variation of life on Earth. Moreover,
they exhibit both modularity and consistency, to the extent that
structures having a particular significance in one location may exhibit
the same significance elsewhere. Obviously, the ultimate
"interpretation" occurs biophysically, in terms of thermodynamics etc,
rather than just by ascription of meaning to arbitrary signifiers, but
there is nevertheless at least an intuitive sense in which the
encoding must obey some structural rules.
As humans, we already have a powerful understanding of rule-based
systems for encoding meaning, because we use them all the time -- I'm
doing so right now as I write, and so are you as you read. Those systems
are languages, and their formal study has given rise to various tools
which might conceivably be applied in a biological context.
Grammars are systems of rules for managing language symbolically,
formally specifying how the symbols -- eg, words -- can legitimately be
strung together and with what structural effects. Grammars operate
primarily at the level of syntax rather than semantics, as this famous
example by Noam Chomsky, the father of modern linguistics, demonstrates:
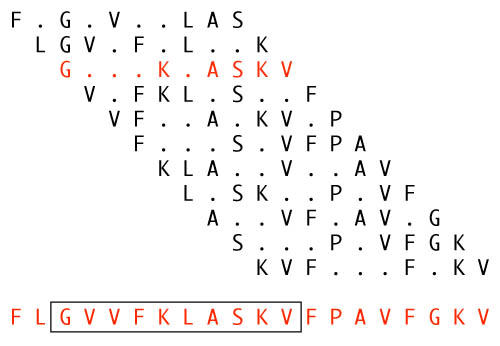
(The 20-mer shown at the bottom is one of the experiment's actual
sequences, D28. The contributing patterns were not published, so those
shown have just been randomly invented to illustrate the concept.)
To avoid too much similarity with real AmPs, sequences with six or more
consecutive amino acids the same as a known AmP sequence were discarded.
Those remaining were ranked according to the popularity of the matched
grammars, and a bunch of promising sequences were synthesised. These
were tested for antimicrobial effects alongside known-good AmPs, known
non-AmPs and other artificial sequences containing the exact same
amino acids but in ungrammatical order.
The results were reasonably promising: nearly half the "designed"
peptides showed antimicrobial effects, and some were competitive with
naturally-occurring AmPs. The shuffled versions, by contrast, were
almost all ineffective, demonstrating the truism that primary structure
is fundamental to protein behaviour.
In this particular case, it appears that even a simplistic language
model has some merit, although it clearly doesn't constitute any kind of
decoding -- it is more like one of those web toys that concoct
movie plots or advertising slogans or whatever by stringing together random words and phrases from
predefined pools in some templated manner:
Colourless green ideas sleep furiously
The sentence is syntactically correct (modulo some reasonable
assumptions as to the grammar we're choosing to parse it with), but
meaningless. This is a distinction that grammars can't, on the whole,
make. (Things, as always, are not quite as clear-cut as this, but let's
ignore that for the moment.)
Chomsky defined a famous hierarchy of grammars that is still widely used
today, according to the difficulty of either generating or parsing the
language. The simplest grammars are Type 3, defining regular
languages, allowing only very basic and mechanical local symbolic
relationships, while the most complex are Type 0, defining
recursively enumerable languages, in which very complex
relationships may exist between widely separated parts of a sentence.
(Type 0 languages are Turing equivalent in the sense that parsing
them requires the same features as universal computation.)
Natural human languages are almost always Type 0, and it is likely that
this is also the general case for biological systems. Nevertheless, many
subsystems may be more tractable, and this paper takes such an approach
with respect to one particular kind of biological sequence.
Antimicrobial peptides are small proteins -- typically of the
order of tens of amino acids in length and without extensive secondary
or tertiary structures -- that occur widely throughout the eukaryote
kingdom as defences against microbial infection. They exploit
differences in cell membrane composition between the host organism and
the microbes to selectively disable or destroy invaders while not posing
a threat to the organism's own cells. These capabilities are static and
comparatively cheap, unlike the more complex responses of an
adaptive immune system; if the latter exists, it will be in
addition to AmP defences rather than as a replacement for them.
One surprising feature of AmPs is that, even though they are very
widespread and have apparently been around in at least some form since
virtually the dawn of time, they seem unsusceptible to resistance.
Possibly their modes of operation are so fundamental that a bacterium
would need to transform itself out of all recognition in order to
escape, with no noticeable selective advantage for the intermediate
states.
Whatever the reason, this offers some hope that AmPs may be a source of
therapies that will remain potent in the face of antibiotic-resistant
agents. However, there is a corresponding concern that modelling drugs
too closely on naturally-occurring AmPs might promote the development of
resistance, with potentially catastrophic consequences for defences on
which we all depend. One suggested avenue is to try to develop
novel AmPs, with similar antimicrobial effects but without close
resemblance to any existing ones.
The authors of this paper hypothesize that AmPs can be modelled
linguistically. Because they are relatively simple, common and very old,
there might be coherent idioms that hold across a wide range of
variations in different organisms. Successful AmPs would operate within
this linguistic space, and our newfangled ones should too, but occupying
portions of the space that have not been significantly colonised so far.
Their experiment is (conceptually) simple: extrapolate AmP idioms to
create novel peptides, then see if they work.
A large number of AmP sequences are known, and these were analysed using
the standard motif-finding program Teiresias. (A
motif is just a shared pattern that is common to the set of
sequences.) The derived motifs were filtered to include only patterns
common to AmPs -- but rare in other non-antimicrobial proteins -- rather
than those common everywhere. The resulting motifs were treated as
regular grammars.
It's important to note a decision made here: that the terminal symbols
of the AmP language will be single amino acid residues. This is
conceptually similar to defining an English grammar in terms of
letters rather than -- as is usual -- parts of speech. For a
human language like English, such an attempt would be doomed to failure.
However, we don't really have any higher-level classification of the
symbolic units of biological sequences that makes any sense in this
context, so there isn't really much choice. Moreover, different amino
acids do have different physiochemical characteristics and could
thus be seen to carry different meanings, so the approach is somewhat
justifiable, despite the overall whiff of desperation.
Having identified about 700 10-residue patterns to be considered
grammatical, 20-residue sequences were generated that conformed to at
least one of those patterns at every constituent 10-residue window, like
this:

Right now, tiaras are plotting to annoint a poor vacuum tube.
My ribcage and lizard are lean, and dentists that I work with may
be beer-stained.
I'm Federal Agent Jack Bauer, and this is the podgiest criminal of my life.
Such an approach clearly has little or no applicability beyond the
particular domain to which it is fitted, and only very constrained
domains are likely to be susceptible to such fitting; nevertheless,
where it works it may serve as a valuable heuristic for pruning a
combinatorially-explosive problem space to the point where workable
solutions can be found.