In our Systems approach to Medicine we bring together Analysis of Medical Data, Modelling biological experiments and Mathematics. From mathematical point of view we have two research directions: modelling complex biological processes, e.g. gene expression or complex molecular machines, and development of new statistical methods to analyse medical data. Our strategic aim is to bring together modelling and data analysis in order to improve public health and suggest new ideas for biotechnology and synthetic biology. To do this we collaborate both with wet biologists and with clinicians. The collaboration structure is displayed here, and you can click on names (under construction) to go to the webpages of our collaborators:
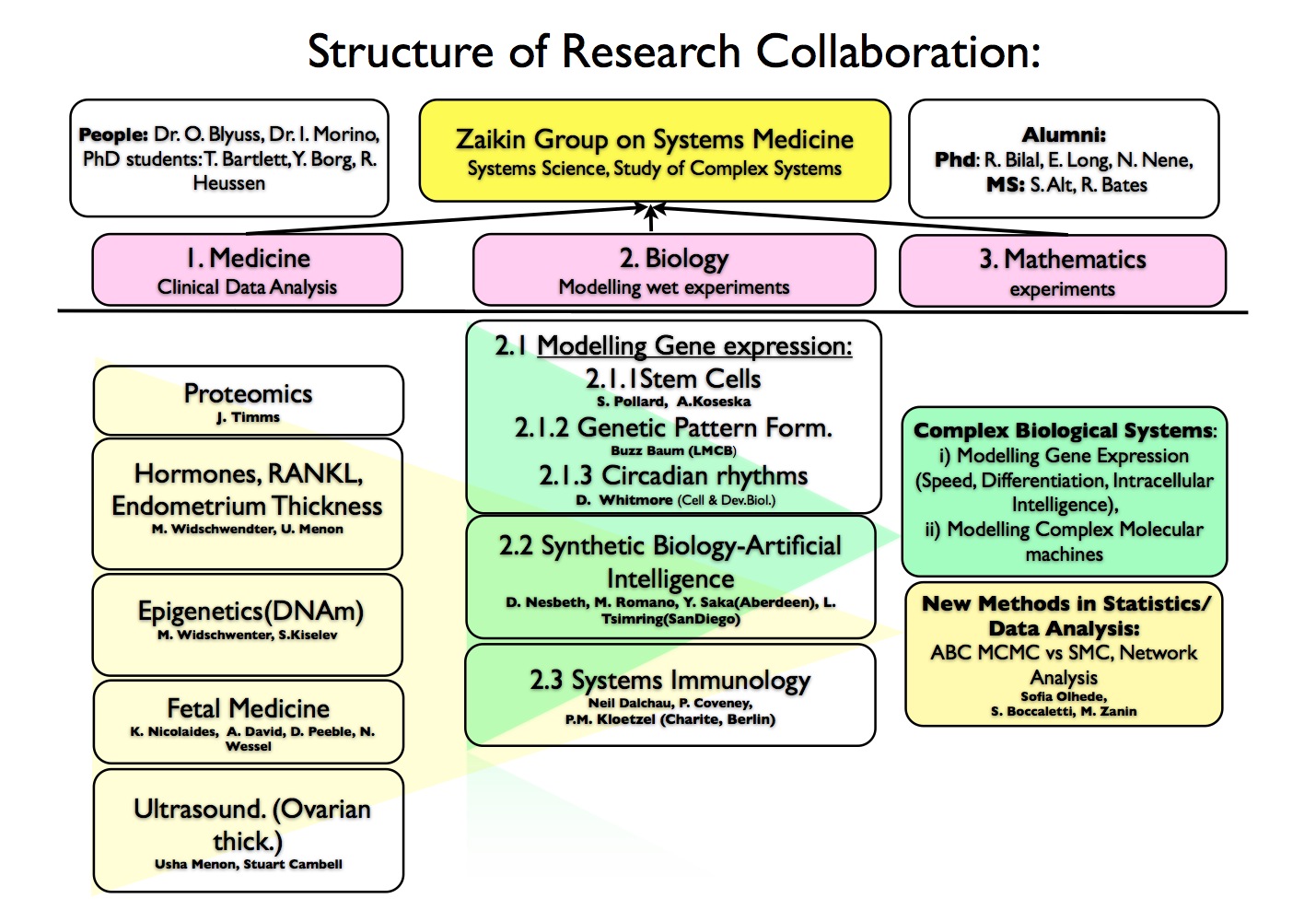
More specifically, at the moment we focus on three research projects:
- Improving early diagnosis and finding new oncomarkers for ovarian and other women cancers based on UKCTOCS screening data. The aim is to find integrated algorithm working with multidimensional serial continuous data (biomarkers) and multidimensional categorical data (genetic and epigenetic risks, epidemiological data). This work is in progress. For initial publications in this direction look for publications 92, 86, 85, 82
- Finding new DNA methylation network measures and new DNA methylation intra-gene measures to improve cancer diagnostics . For initial publications in this direction look for publications 89, 84, 95
- Investigation of complex genetic dynamics in genetic networks, with applications to Synthetic Biology, Circadian Oscillators, Stem Cell differentiation and genetic pattern formation. We have found and interested in several new effects or genetic systems: distributed genetic classifiers (97), intelligent and noisy genetic perceptrons (91, 98), speed dependent cellular decision making (78, 80, 83), timed cellular decision making vis cell-cell communication (70), multistability and clustering of genetic oscillators (63, 62, 69, 60), quantized time production cycling time in gene networks (59).
We are still interested in our previous research projects:
- Modelling complex molecular machines - proteasomes, and finding new methods to detect proteasomal splicing (73, 67, 68, 71, 58, 57).
- Modelling complex bone dynamics (66, 48, 49).
- Investigating new noise induced effects: doubly stochastic effects (34), doubly stochastic resonance (22), vibrational resonance (43, 42), oscillatory amplification of stochastic resonance in excitable systems (40), doubly stochastic coherence (37), noise-induced excitability (36), systems size resonance (35), noise-induced propagation (32), and noise-induced transitions (24).
We provide methodological support for medical groups in Institute for Women's Health and develop new statistical methods, especially in:
- Experimental design
- Descriptive statsictics: multivariate logistics regression, odds ratios and statistical testing
- Machine learning algorithms: clustering, SVM
- Bayesian methods and Markov Chain Monte Carlo Methods, e.g., Bayesian models with changing points
- Network and graph-based statistical methods