back to B250 timetable back to Hopeful Monsters back to segmentation
What is a regulatory mutation?
All cells in an organism contain the same genome - that is, the same genes arranged in the same order along the same chromosomes. This is a generalization - some genes are rearranged within certain cells, but this is the exception rather than the rule. For example, the genes coding for immunoglobulins (antibody molecules) are rearranged in B lymphocytes; this is how novel antibody structures are created in response to new invading organisms or viruses. Also, chromosomes are often broken and rejoined aberrantly in cancer cells. And, of course, germ cells (sperm and eggs) contain only half the number of chromosomes as normal somatic (body) cells.
Nevertheless, most cells contain identical DNA. What makes one type of cell different from another is not the genes they contain but which of these genes they express - i.e. which genes they transcribe into RNA, then translate into protein. A muscle cell differs from a neuron in a great many of its constituent RNAs and proteins. For example, the muscle cell transcribes the genes coding for muscle actin and myosin but not the gene encoding neurofilament; the reverse is true of the neuronal cell. What determines whether a particular gene is transcribed or not? This depends to a large extent on the other proteins that are present in the cell - in particular, proteins that are collectively known as "transcription factors".
Every gene consists of a protein coding sequence, which might be contiguous or broken up into a series of exons and introns, and which begins with a START codon (ATG) and concludes with a STOP codon (TAA, TAG or TGA). Apart from this, a gene must have regulatory sequences associated with it. These are stretches of DNA which do not themselves code for protein but which act as binding sites for RNA polymerase and its accessory molecules as well as a variety of transcription factors. Together, the regulatory sequences with their bound proteins act as molecular switches that determine the activity state of the gene - e.g. OFF or FULL-ON or, more often, something in between. The regulatory sequences include the promoter region together with enhancer elements.
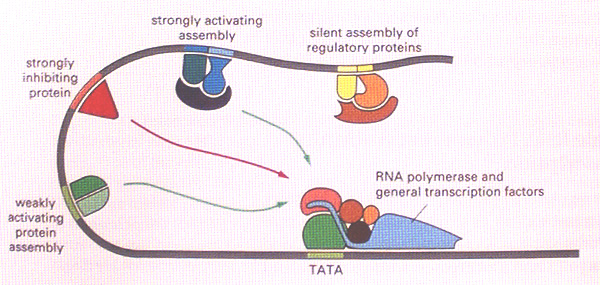
Every gene has a promoter, which is the binding site for the basal transcriptional apparatus - RNA polymerase and its co-factors. This provides the minimum machinery necessary to allow transcription of the gene. The enhancer regions are found at a distance from the promoter, to either the5' or 3' sides of the gene or within introns. They are typically short stretches of DNA (200bp, say), each made up of a cluster of even shorter sequences (25bp, say) that are the binding sites for a variety of cell- or region-specific transcription factors. Once bound, these transcription factor complexes interact with the basal transcriptional machinery at the promoter to enhance (or sometimes diminish) the transcription rate of the gene. Such interactions are possible because of the flexible nature of DNA which allows the enhancers to come close to the promoter by looping out the DNA in between (see diagram below).
We can think of the activating function of enhancers as follows. Binding of RNA polymerase and the basal transcriptional machinery at the gene promoter is like switching on the engine and allowing it to idle in neutral. When the supplementary transcription factors bound to enhancer elements interact with the basal machinery, it is like putting the engine into gear and pulling away from the kerb. (Alternatively, for a repressor site it is like putting on the handbrake.)
Frequently, a given gene is subject to complex regulation. That is, it might have to be transcribed at different times and in different places during development, or in response to different extracellular stimuli. In the present context (Drosophila embryogenesis) we have seen that the segmentation genes are expressed according to their position in the embryo. An example is the even-skipped (eve) gene, a pair-rule gene that is transcribed in alternate embryonic parasegments to generate a zebra pattern of seven stripes. The transcriptional state of the eve gene - either ON or OFF according to which parasegment we are in - is under the control of a series of enhancer elements, one for each stripe. Each enhancer element contains binding sites for upstream segmentation gene products such as Bicoid and Kruppel (which, as you will recall, are themselves transcription factors). Thus the particular constellation of maternal effect genes, gap genes and other pair-rule genes that is expressed in a given parasegment determines whether or not one of the enhancer elements is fully occupied and consequently whether eve gene transcription is activated or not in that parasegment. The specificity of the enhancers can be demonstrated by removing just one of them (the element that specifies stripe 2, say) and inserting it upstream of a reporter gene like the bacterial beta-galactosidase (Bgal). when this is introduced into the embryo, Bgal is expressed in just one stripe - in the position of the eve stripe 2. Alternatively, particular enhancer elements can be deleted from the normal eve gene, resulting in deletion of the corresponding stripes of eve expression. Such a mutation - loss of an enhancer element - is called a regulatory mutation. It affects the spatial or temporal regulation of the gene without causing universal loss of the gene product.
 The upper part of this diagram shows part of the regulatory
region of the eve gene - the part closest to the transcription start
site (red arrow). In this region there are three enhancer elements that control
eve expression in stripes 2, 3 and 7. Other elements, not shown in the
diagram, lie further away from the transcription start site (to the left) and
control eve expression in stripes 1,4,5 and 6. Deleting these upstream
elements and leaving only those shown in the diagram gives rise to the
expression pattern shown above right - stripes 2, 3 and 7. Taking just one of
the elements, that for stripe 2, and linking it to a reporter gene gives the
pattern shown above centre - stripe 2 only. The wild-type eve
expression pattern is shown above left. (From Gilbert, Developmental Biology
4th edition Chapter 15, p549 and Alberts et al. Molecular Biology of the Cell
3rd edition, Chapter 9 p428)
The upper part of this diagram shows part of the regulatory
region of the eve gene - the part closest to the transcription start
site (red arrow). In this region there are three enhancer elements that control
eve expression in stripes 2, 3 and 7. Other elements, not shown in the
diagram, lie further away from the transcription start site (to the left) and
control eve expression in stripes 1,4,5 and 6. Deleting these upstream
elements and leaving only those shown in the diagram gives rise to the
expression pattern shown above right - stripes 2, 3 and 7. Taking just one of
the elements, that for stripe 2, and linking it to a reporter gene gives the
pattern shown above centre - stripe 2 only. The wild-type eve
expression pattern is shown above left. (From Gilbert, Developmental Biology
4th edition Chapter 15, p549 and Alberts et al. Molecular Biology of the Cell
3rd edition, Chapter 9 p428)
A similar thing goes for regulation of the Bithorax Complex. Transcription of Ubx, abd-A and Abd-B is controlled by a series of enhancer elements, each of which is specific for a particular parasegment. Loss of one of the Ubx enhancers will result in loss of Ubx expression from the corresponding parasegment and transformation of that parasegment. The BX-C mutations picked up by Lewis in his genetic screen (bx, pbx, bxd, iab2 etc) turn out to be regulatory mutations of this sort. For example, bxd controls Ubx expression in A1; mutations of bxd result in loss of Ubx expression in A1 and homeotic transformation of A1 to T3). Thus, transformations of segment identity result from subtle, parasegment-specific alterations to the expression patterns of Ubx, abd-A or Abd-B, not complete loss of their encoded proteins.
back to B250 timetable back to Hopeful Monsters back to segmentation