Analysis of human serum proteomes for cancer biomarker discovery
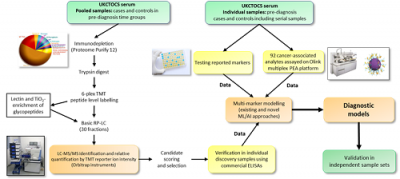
By using samples predating diagnosis we hope to identify early markers in the absence of non-specific late stage changes.
Biomarker discovery work uses multi-dimensional protein and peptide separations linked to tandem MS with isobaric mass tagging for quantification.
By comparing profiles of serum from healthy volunteers, patients with benign and malignant disease our goal is to identify candidate biomarkers for differential diagnosis and early detection of ovarian, breast, pancreaticobiliary and colorectal cancers and endometriosis.
Promising markers are being further validated in independent sample sets using immune- and MS-based assays and multi-marker models developed for translation to the clinic.
ErbB2/HER2 signalling in breast cancer
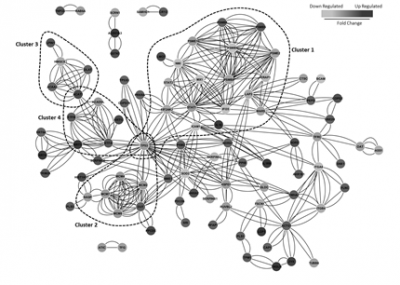
Parallel differential mRNA and protein expression analyses have been applied to profile model breast cancer cell lines to understand how target genes are regulated by ErbB2 overexpression and specific receptor ligands.
Targets of interest are being validated and functionally characterised using cell-based assays following siRNA-mediated knockdown.
Further work applying protein expression and phosphoproteomic profiling methods is aimed at understanding ErbB signalling and regulated expression at the molecular level and relating this to cellular phenotype
Ovarian cancer biology
This research is aimed at a better understanding of the molecular events required for ovarian cell carcinogenesis, tumour progression and chemoresistance, and to thereby identify potential diagnostic, prognostic and predictive biomarkers for ovarian cancer.
Work involves the application of complementary proteomic technologies to profile novel cell models of early ovarian epithelial cell transformation, metastasis, tumour suppression and chemoresistance.
Candidate genes of interest are further validated and functionally characterised using RNAi-mediated knockdown and assessment of phenotype using cell-based assays.
 Close
Close

