New publication in Developmental Cell for Mao Lab
Multiple tissues in our body, such as the brain, and the gut, are folded structures. Similar to an origami crane emerging from the folds of a paper, all growing tissues must fold in precise positions, at multiple steps during development, to reach the correct 3D adult shape. If these folds do not form in the correct way, the structure and function of the organ will be disrupted. A new publication in Developmental Cell for the Mao Lab explains how this precise positioning of the folds can be achieved by controlling the growth patterns of the tissue. They have created a new computer model, a virtual tissue growth platform, which simulates how tissues grow and fold, much like a computer game. In their computer model, they can predict the shapes of normal and mutant animal tissues. With this model, they have the potential now to predict the sizes and shapes of multiple normal and diseased organs, and assist doctors in designing medical interventions, as well as in guiding tissue engineers in growing artificial organs for regenerative medicine.
Abstract from the article:
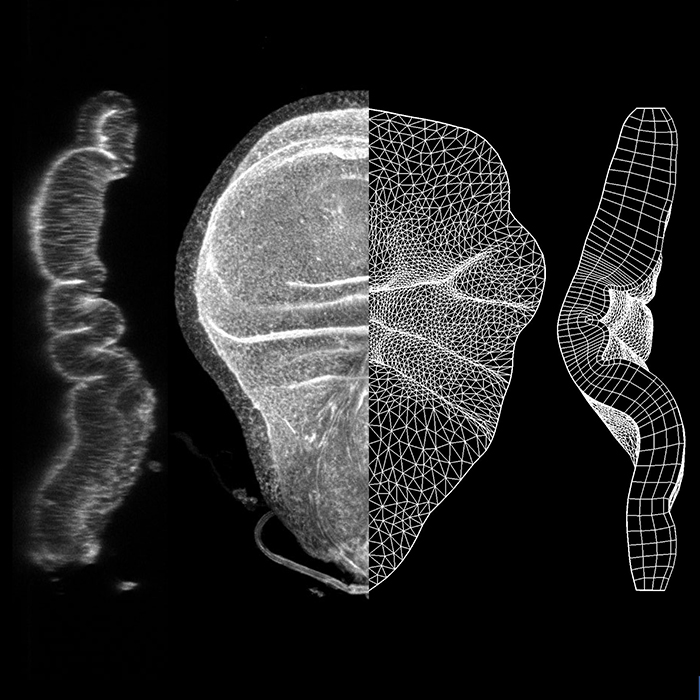
Tissue folding is a fundamental process that shapes epithelia into complex 3D organs. The initial positioning of folds is the foundation for the emergence of correct tissue morphology. Mechanisms forming individual folds have been studied, but the precise positioning of folds in complex, multi-folded epithelia is less well-understood. We present a computational model of morphogenesis, encompassing local differential growth and tissue mechanics, to investigate tissue fold positioning. We use the Drosophila wing disc as our model system and show that there is spatial-temporal heterogeneity in its planar growth rates. This differential growth, especially at the early stages of development, is the main driver for fold positioning. Increased apical layer stiffness and confinement by the basement membrane drive fold formation but influence positioning to a lesser degree. The model successfully predicts the in vivo morphology of overgrowth clones and wingless mutants via perturbations solely on planar differential growth in silico.
 Close
Close