Our current research is diverse and interdisciplinary - including arsenic metabolism, bioleaching and extremophiles, and bacteriophages and gut microbiomes.
- Arsenic Metabolism
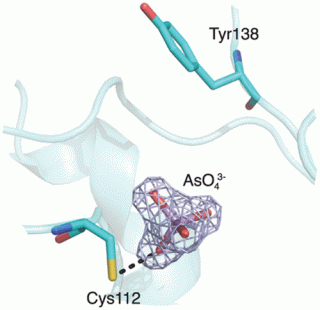
Two soluble forms of arsenic are found in nature: arsenite [As(III)] and arsenate [As(V)]. Both of these forms are toxic and represent a threat to our health (e.g., in Bangladesh and West Bengal) and environment (owing often to human activities such as mining).
Microbes can use toxic substances such as arsenic for growth, by using arsenate as an electron acceptor (i.e., for respiration) or arsenite as an electron donor (with either oxygen or nitrate as an electron acceptor).
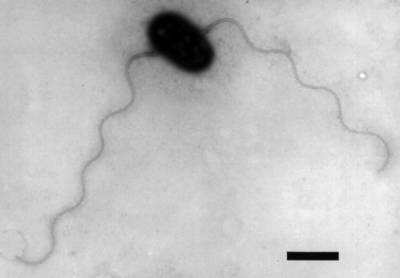
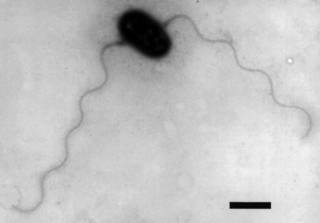
Arsenite-oxidising bacterium NT-26

Hand pump from arsenic-contaminated tube well in Bangladesh
Research Questions:
- how do microbes use arsenic for growth (i.e. what is the mechanism)?
- how is the metabolism regulated?
- what is the distribution and abundance of the genes that encode these properties, in various environments?
- what is the role of these organisms in the cycling of arsenic in the environment?
- what is the origin of the enzymes involved in the metabolism?
- how can we use the metabolic enzymes for biosensors or organisms for bioremediation?
- Bacteriophages
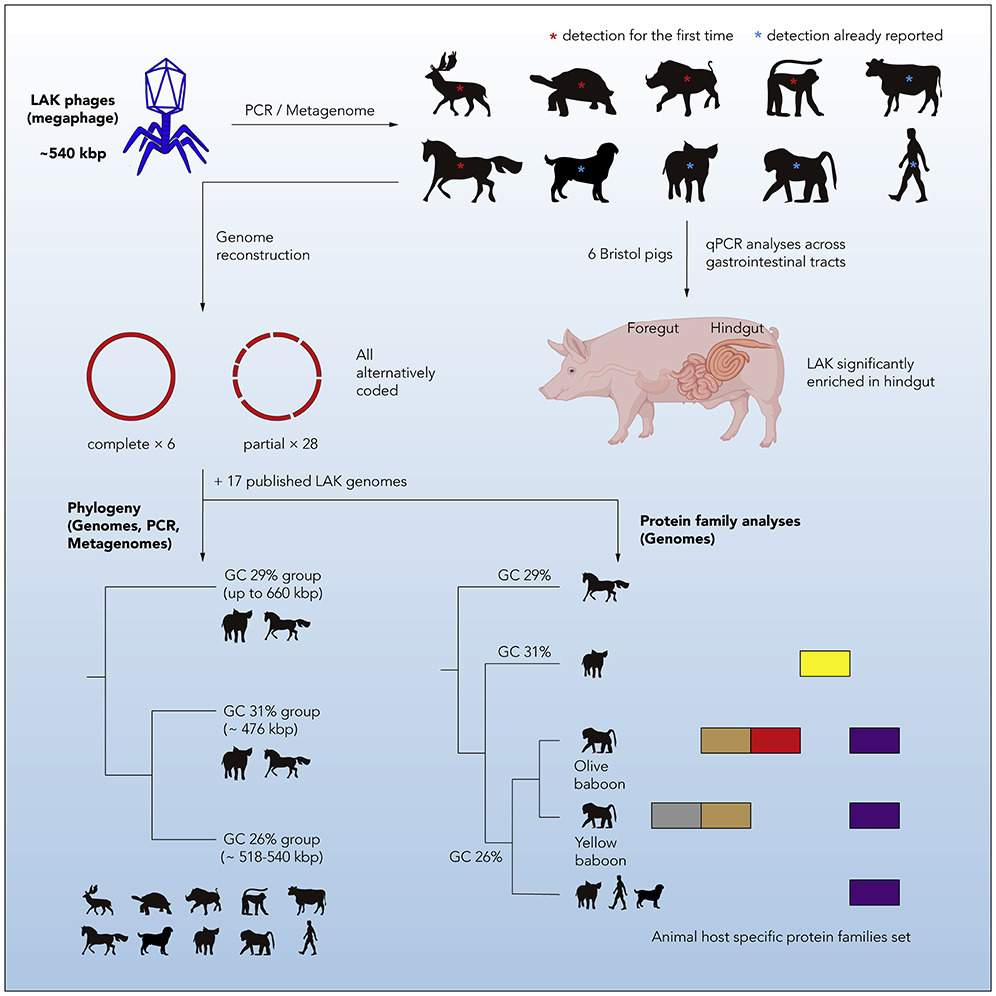
Bacteriophages (phages; viruses of bacteria) are the most abundant organisms on earth. Despite this, phages are largely overlooked components of the gut microbiome. Gut phages have the capacity for nutrient redistribution through specific bacterial lysis. Furthermore, prophages (latent phage) are able to induce bacterial horizontal gene transfer to facilitate host pathogenic evolution. Together, this can shape bacterial community structure.
Genome-resolved meta-genomics conducted by our collaborators at UC Berkley uncovered Lak phages in the gut microbiome of Bangladeshi adults. These novel ‘Mega phages’ (with genomes >540 kb) prey on bacteria of the genus Prevotella; which play a role in fibre degradation.
Our work has since confirmed the presence of Lak in different human cohorts, baboons, pigs and other animals consuming high-fibre diets.
Research Objectives:
- Uncovering and exploring the genetic diversity of Lak phage variants found in different animals
- Studying the dynamics between phage and bacterial host in the gut
- Screening samples for novel lytic bacteriophages with potentially therapeutic and/or undesirable outcomes
- Bioleaching
Bioleaching is a cost-effective, low input way of extracting precious metals from sulfide minerals using bacteria and archaea collected from acid mine environments. The process exploits the sulfur and iron metabolisms of these microbes to break down ore, allowing the target metals to be obtained. Our research uses molecular biology and bioinformatic techniques to explore what roles different microbial community members are playing during the progression of bioleaching. The results of this work could help refine the bioleaching process, making metal mining more efficient and environmentally-friendly.
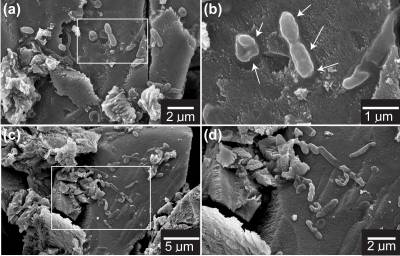
Microbial consortium on the mineral chalcoyrite

Copper mine in Cyprus
Research Questions:
- What interactions do extremophiles have with sulfide minerals?
- What are the community dynamics of bioleaching consortia?
- What roles do different microbial community members play as minerals breakdown during bioleaching?
- What genes associated with sulfur and iron metabolisms are different community members expressing as bioleaching progresses?
 Close
Close