|
Daniel Manson
PhD student, Institute of Cognitive Neuroscience Alexandra House, 17 Queen Square, UCL London WC1N 3AR Doctoral training centre CoMPLEX: Centre for Mathematics and Physics in the Life Sciences and Experimental Biology ucl.ac.uk/complex | UCL Physics Building, Gower Street | +44 (0)20 7679 4325 | complex@ucl.ac.uk  |
|
|
About
Life before UCL. I was born and raised in London, and graduated from Warwick University in 2009 with a BSc in Mathematics and Physics. I then spent a year as an English teacher in Moscow, where I realised that I wanted to do a PhD in Neuroscience.At UCL. In 2010-11 I worked as a postgraduate researcher in Neil Burgess's lab at UCL's Institute of Cognitive Neuroscience. The lab investigates spatial representation in the brain, primarily using extracellular recording of rat hippocampus and entorhinal cortex. Much of the work I did was data analysis, but I did also undertake a small amount of experimental work. Neil's postdoc Caswell Barry provided day-to-day supervision.
In October 2011 I joined UCL's CoMPLEX doctoral training centre on its MRes + PhD program. In the first few weeks of the program I received a formal introduction to various aspects of cutting edge biology. I then undertook a series of three six-week projects followed by a longer project during the summer. In October 2012 I returned to Neil Burgess's lab, now as a PhD student co-supervised by John O'Keefe (winner of the 2014 Nobel Prize for Physiology or Medicine).
show/hide abstracts
Publications
Using Grid Cells for NavigationNeuron, 2015 | Neuron link | D. Bush, C. Barry, D. Manson, N. Burgess
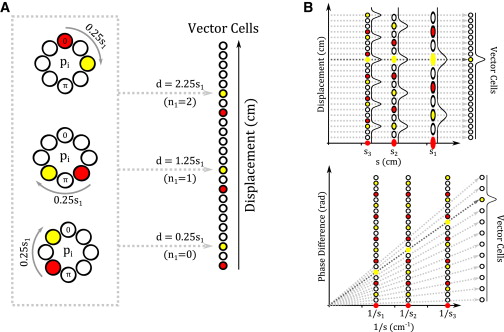 Mammals are able to navigate to hidden goal locations by direct routes that may traverse previously unvisited terrain. Empirical evidence suggests that this “vector navigation” relies on an internal representation of space provided by the hippocampal formation. The periodic spatial firing patterns of grid cells in the hippocampal formation offer a compact combinatorial code for location within large-scale space. Here, we consider the computational problem of how to determine the vector between start and goal locations encoded by the firing of grid cells when this vector may be much longer than the largest grid scale. First, we present an algorithmic solution to the problem, inspired by the Fourier shift theorem. Second, we describe several potential neural network implementations of this solution that combine efficiency of search and biological plausibility. Finally, we discuss the empirical predictions of these implementations and their relationship to the anatomy and electrophysiology of the hippocampal formation.
Mammals are able to navigate to hidden goal locations by direct routes that may traverse previously unvisited terrain. Empirical evidence suggests that this “vector navigation” relies on an internal representation of space provided by the hippocampal formation. The periodic spatial firing patterns of grid cells in the hippocampal formation offer a compact combinatorial code for location within large-scale space. Here, we consider the computational problem of how to determine the vector between start and goal locations encoded by the firing of grid cells when this vector may be much longer than the largest grid scale. First, we present an algorithmic solution to the problem, inspired by the Fourier shift theorem. Second, we describe several potential neural network implementations of this solution that combine efficiency of search and biological plausibility. Finally, we discuss the empirical predictions of these implementations and their relationship to the anatomy and electrophysiology of the hippocampal formation.
Grid Cells Form a Global Representation of Connected Environments
Current Biology, 2015 | ScienceDirect link | F. Carpenter, D. Manson, K. Jeffery, N. Burgess, C. Barry
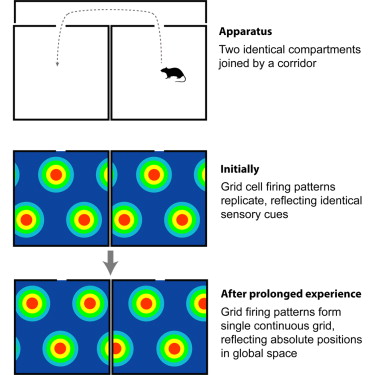 The firing patterns of grid cells in medial entorhinal cortex (mEC) and associated brain areas form triangular arrays that tessellate the environment and maintain constant spatial offsets to each other between environments. These cells are thought to provide an efficient metric for navigation in large-scale space. However, an accurate and universal metric requires grid cell firing patterns to uniformly cover the space to be navigated, in contrast to recent demonstrations that environmental features such as boundaries can distort and fragment grid patterns. To establish whether grid firing is determined by local environmental cues, or provides a coherent global representation, we recorded mEC grid cells in rats foraging in an environment containing two perceptually identical compartments connected via a corridor. During initial exposures to the multicompartment environment, grid firing patterns were dominated by local environmental cues, replicating between the two compartments. However, with prolonged experience, grid cell firing patterns formed a single, continuous representation that spanned both compartments. Thus, we provide the first evidence that in a complex environment, grid cell firing can form the coherent global pattern necessary for them to act as a metric capable of supporting large-scale spatial navigation.
The firing patterns of grid cells in medial entorhinal cortex (mEC) and associated brain areas form triangular arrays that tessellate the environment and maintain constant spatial offsets to each other between environments. These cells are thought to provide an efficient metric for navigation in large-scale space. However, an accurate and universal metric requires grid cell firing patterns to uniformly cover the space to be navigated, in contrast to recent demonstrations that environmental features such as boundaries can distort and fragment grid patterns. To establish whether grid firing is determined by local environmental cues, or provides a coherent global representation, we recorded mEC grid cells in rats foraging in an environment containing two perceptually identical compartments connected via a corridor. During initial exposures to the multicompartment environment, grid firing patterns were dominated by local environmental cues, replicating between the two compartments. However, with prolonged experience, grid cell firing patterns formed a single, continuous representation that spanned both compartments. Thus, we provide the first evidence that in a complex environment, grid cell firing can form the coherent global pattern necessary for them to act as a metric capable of supporting large-scale spatial navigation.
The scale of grid cell firing patterns is modulated by spatial uncertainty [Poster]
SfN, 2014 | SfN abstract page | D. Manson, F. Carpenter, J. O'Keefe, N. Burgess, C. Barry
The representation of self-location by grid cells has variously been described as “densely packed” or “hexagonal”; leading to hypotheses about the role of a grid cell network in providing a metric for space. However, recently, it was shown that exposure to environmental novelty provokes a temporary increase in grid scale, which attenuates with increasing familiarity. This differs from evidence that grids are anchored to sensory cues, and can be rotated or stretched by changes to sensory cues, and suggests that grid cells do not simply provide a fixed metric for space that becomes attached to environmental cues. The exact neural mechanisms driving grid scale expansion and its functional role are currently unknown. However, it has been hypothesised that grid scale varies inversely with the precision with which the animal’s self-location can be determined (spatial uncertainty), so as to encode location more accurately. Thus grid scale would be expected to increase in situations with few spatial cues or unknown configurations of cues, such as in a novel environment, compared to situations in which multiple cues were accessible. To test this hypothesis, we recorded medial entorhinal grid cells while rats foraged in two types of environment: one rich in local cues and the other lacking any such cues. It was hypothesised that grid scale would be smaller in the cue rich environment, reflecting higher certainty in the representation of self-location, similar to that observed in familiar environments. We recorded cells in both environments from their first experience of it and throughout the familiarisation process. We found that grid patterns had a smaller scale in the cue rich environment, supporting the hypothesis, and were also more regular. This confirms that the grid cells do not simply encode a fixed metric for space, rather the encoding is sensitive to additional factors such as environmental novelty and the availability of cues.
Theta phase precession of grid and place cell firing in open environments.
Philos Trans R Soc B, 2013 | PubMed link | A. Jeewajee, C. Barry, V. Douchamps, D. Manson, C. Lever, N. Burgess
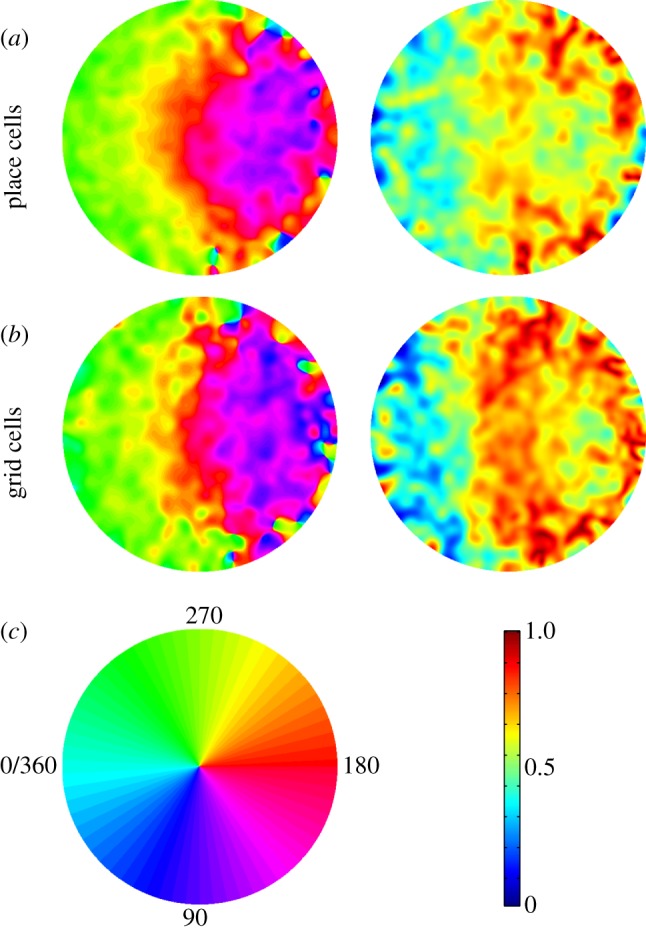 Place and grid cells in the rodent hippocampal formation tend to fire spikes at successively earlier phases relative to the local field potential theta rhythm as the animal runs through the cell's firing field on a linear track. However, this 'phase precession' effect is less well characterized during foraging in two-dimensional open field environments. Here, we mapped runs through the firing fields onto a unit circle to pool data from multiple runs. We asked which of seven behavioural and physiological variables show the best circular-linear correlation with the theta phase of spikes from place cells in hippocampal area CA1 and from grid cells from superficial layers of medial entorhinal cortex. The best correlate was the distance to the firing field peak projected onto the animal's current running direction. This was significantly stronger than other correlates, such as instantaneous firing rate and time-in-field, but similar in strength to correlates with other measures of distance travelled through the firing field. Phase precession was stronger in place cells than grid cells overall, and robust phase precession was seen in traversals through firing field peripheries (although somewhat less than in traversals through the centre), consistent with phase coding of displacement along the current direction. This type of phase coding, of place field distance ahead of or behind the animal, may be useful for allowing calculation of goal directions during navigation.
Place and grid cells in the rodent hippocampal formation tend to fire spikes at successively earlier phases relative to the local field potential theta rhythm as the animal runs through the cell's firing field on a linear track. However, this 'phase precession' effect is less well characterized during foraging in two-dimensional open field environments. Here, we mapped runs through the firing fields onto a unit circle to pool data from multiple runs. We asked which of seven behavioural and physiological variables show the best circular-linear correlation with the theta phase of spikes from place cells in hippocampal area CA1 and from grid cells from superficial layers of medial entorhinal cortex. The best correlate was the distance to the firing field peak projected onto the animal's current running direction. This was significantly stronger than other correlates, such as instantaneous firing rate and time-in-field, but similar in strength to correlates with other measures of distance travelled through the firing field. Phase precession was stronger in place cells than grid cells overall, and robust phase precession was seen in traversals through firing field peripheries (although somewhat less than in traversals through the centre), consistent with phase coding of displacement along the current direction. This type of phase coding, of place field distance ahead of or behind the animal, may be useful for allowing calculation of goal directions during navigation. show/hide abstracts
MRes work
Contrast Invariance of Cell Populations in V1Jan 2012 | pdf | supervised by Matteo Carandini
Cells in mammalian primary visual cortex respond preferentially to bars of lightness or darkness on the retina. For a long time it has been known that the extent of this sensitivity does not depend on the contrast of the stimulus. It has recently been shown that this invariance also exists in the shape of the time-averaged population response. Here we examine whether invariance of the population holds over much shorter time periods, and tentatively conclude that it does.
Modeling the Metabolic Interactions of Astrocytes and Neurons
Under Normal Conditions and During Ischemic Hypoxia
Mar 2012 | pdf | supervised by Ilias Tachtsidis
It is relatively common for the brain of perinatal infants to be exposed to a period of reduced oxygen (hypoxia) and reduced blood flow (ischemia). Left untreated, these episodes tend to cause permanent damage to the brain. In order to understand how this damage arises and whether it is preventable, an extensive computational model has been developed that attempts to accurately capture the full metabolic complexity of the infant brain. This complexity encompasses everything from haemodynamics, to calcium channels in vascular muscle, to oxidative phosphorylation in the mitochondria of neurons. Model parameters are set using data from in vivo and in vitro experiments, as well as using a range of more ad hoc measures such as steady-state analysis. In the current model there is no distinction between astrocytes and neurons, i.e. the cell in the model is a rough average of the two. However there is plenty of evidence that astrocytes and neurons have quite different metabolic properties and interact in non-trivial ways. In this work we examine a separate model, designed for looking at the metabolic interactions of astrocytes and neurons, and examine whether any of its dynamics should be incorporated into the hypoxia model.
Endoscopic Transplantation of Esophageal Mucosa Grown In Vitro
Using Magnetic Particles in an Externally Administered Magnetic Field
May 2012 | pdf | supervised by Richard Day and Quentin Pankhurst
If identified at an early stage, cancer of the esophagus can be removed non-invasively using an endoscope. However, the site of removed tissue rarely recovers completely, and is commonly replaced by a large volume of scar tissue, in some cases causing a significant narrowing of the esophagus. To prevent such post-operative problems it has been proposed that healthy replacement tissue be applied to the site of the cancer, having been grown in vitro from the patient's own cells. In this work we consider one method for improving the adhesion of the new cells to the target site. The method involves seeding the in vitro cells with magnetic particles, and subsequently applying a magnetic force to them using an externally administered permanent magnet, applied for a few minutes during the grafting procedure.
The Seas are Full of Krill (Poster)
Jun 2012 | pdf | transferable skills mini-project
Following a week-long course at the Marine Biology Assosiation's laboratory in Plymouth, MRes students are required to produce a poster relating to something they have learnt. Producing and presenting the poster gives students a chance to practice important 'transferable' skills. I chose to make a poster on krill and their role in the marine carbon system.
Characterizing Brain Activity at the Mesoscopic Scale
Comparing a Kuramoto model to Resting State MEG data
August 2012 | pdf | supervised by Luc Berthouze and Simon Farmer
Despite impressive advances over the last several decades, to a large extent the human brain remains something of a black-box. This state of affairs is easily understood given that the `machine inside the box' consists of a network of approximately 1011 nodes, each of which transmits electrical signals in a highly non-linear and ever changing manner. Investigation of individual cell types has been relatively successful in primary sensorary areas and hippocompus, but there is yet
to be any signiffcant progress in understanding the dynamics of small populations of cells. Appropriate grouping of these small populations results in a brain divided into approximately 1000 regions. These regions can then be further aggregated to produce a labeled map of the brain at the mesoscopic scale where there are about 66 distinct regions. EEG, MEG and fMRI make it possible to record brain activity at this coarse level of network description. In this work we model the 66 regions as a network of weakly coupled oscillators, i.e as a Kuramoto model, and compare model simulations to MEG data collected at resting state.
Interests
I am a keen programmer. For a couple of years I was a big fan of MATLAB, but now consider myself a Python evangelist. I also dabble in C/C++ and various HTML5 bits and bobs (the web browser is a pretty powerful tool when it comes to making highly interactive user interfaces).Pretty much all aspects of neuroscience interest me: from the philosophical (Dennett's pretty good) to the biochemical, and everything in between. The algorithmic/cognitive/systems level of research I find most fascinating (you can't beat Marr).
I hope that one day we will have the tools to run the brain 'in debug mode' and really find out what is going on. In the meantime we shall have to content ourselves with staring at the "synthetic brains" of Deep Neural Networks.
