Structural and Molecular Biology
We are the Research Department of discoveries, new ideas and methodologies in biomolecular sciences. Within the stimulating and diverse research and training environment of international standing, we lead and advance on essential questions in exciting areas of modern biochemistry, biotechnology, and medicine at the atomic, molecular, cellular and organism levels.
Owing to the strength and the technical expertise of its research community SMB is a key player in many collaborative multi-disciplinary projects across UCL, nationally and internationally. This vision is enhanced via our partnership with the Department of Biological Sciences at Birkbeck College and the Institute of Structural and Molecular Biology (ISMB).
Emerging from the rich history of Biochemistry Departments at UCL, nowadays, the SMB provides a home to engaged, enthusiastic and well-supported staff and students. We learn and teach through our research; we nurture curiosity, welcome challenging questions, and embrace all voices.
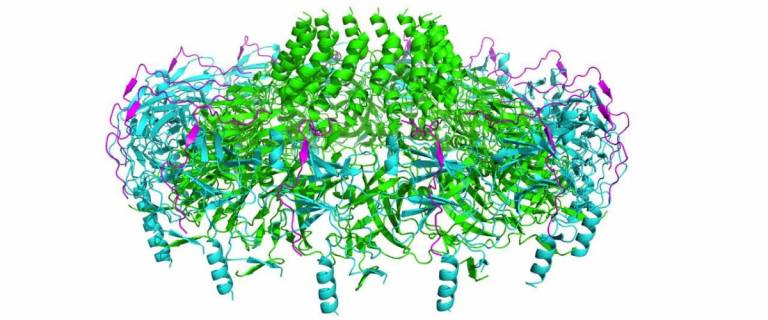
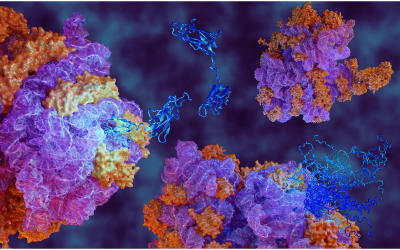
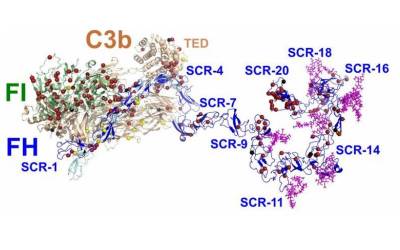
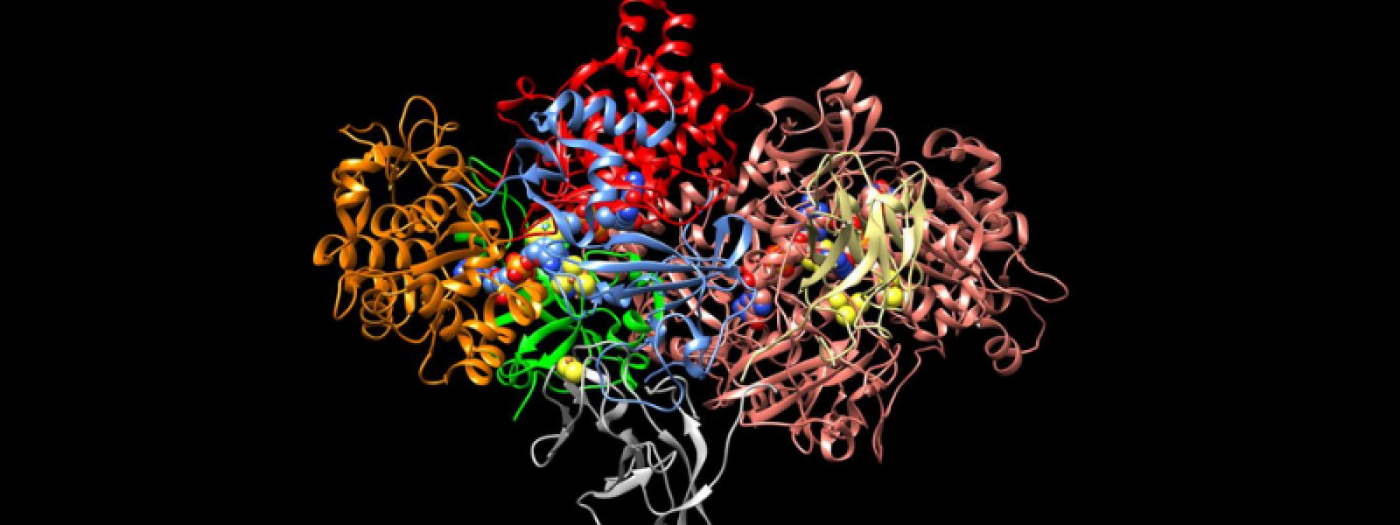
Atomic resolution structure of the core complex of a type IV secretion system involved in bacterial conjugation (Nature (2009) 462:2011-2015)
Our Department
Research
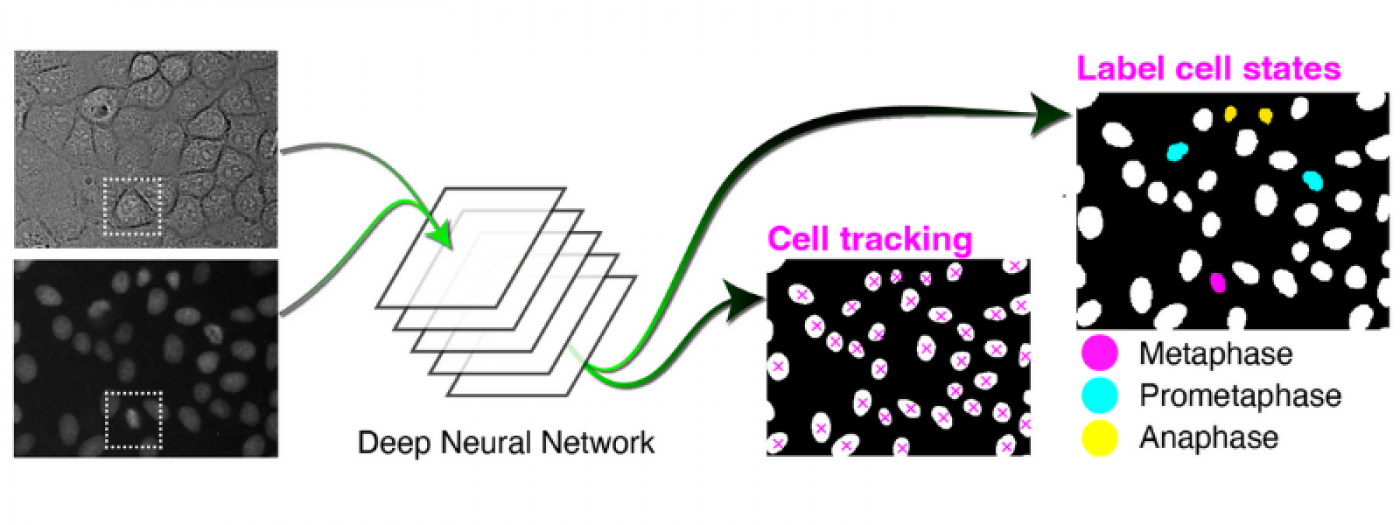
Computational Biology and AI
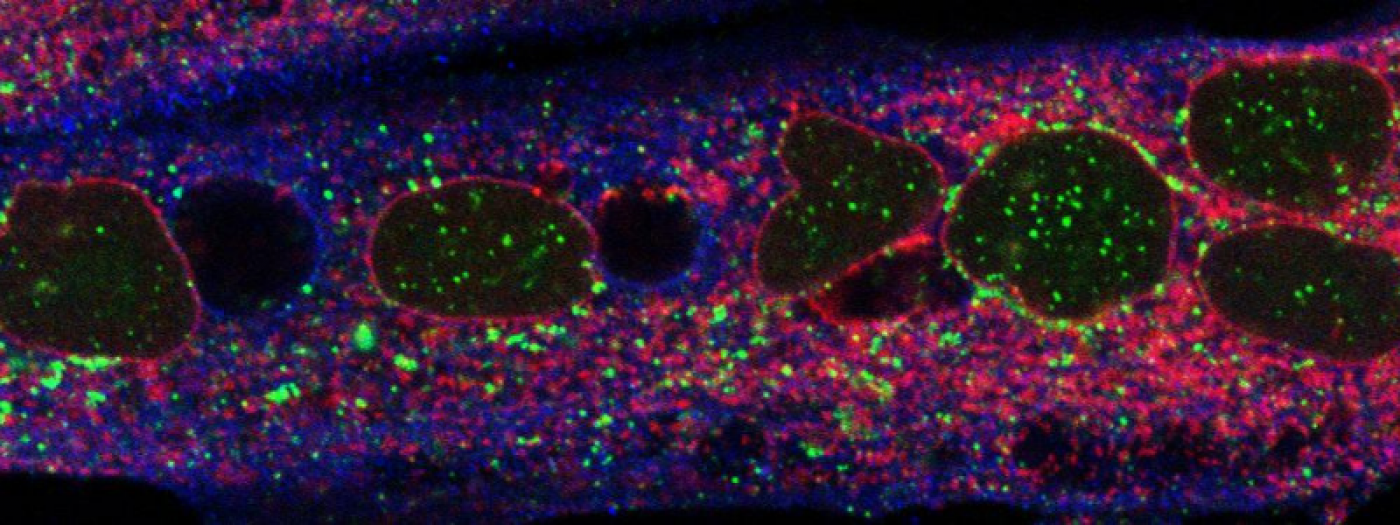
Membranes, Signalling and Trafficking
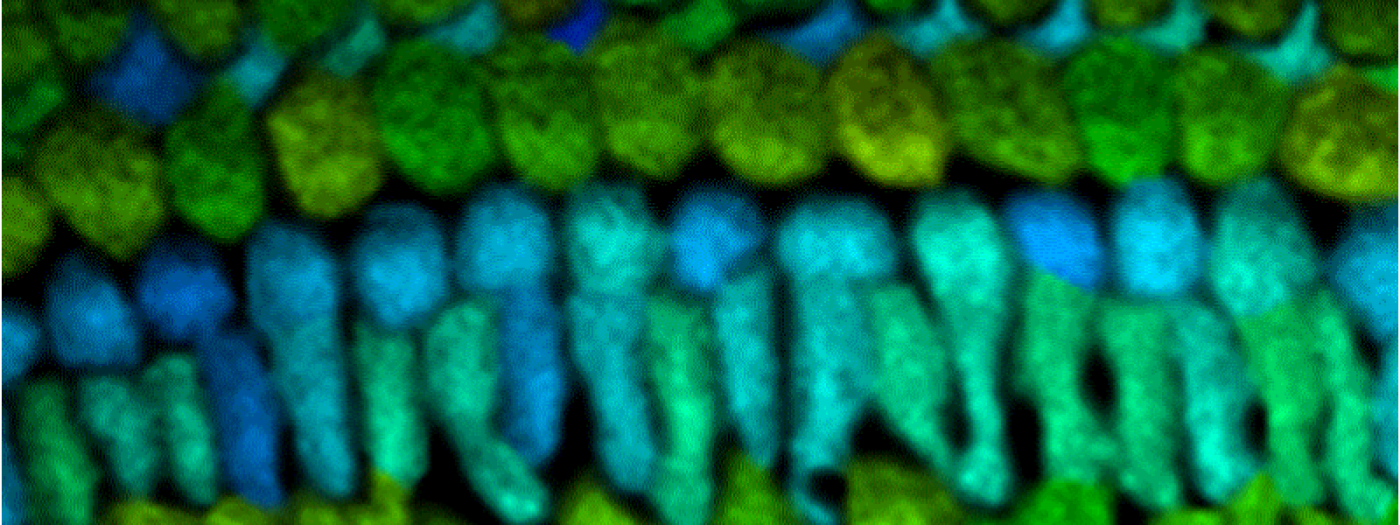
Metabolism and Disease
Microbes and Environment
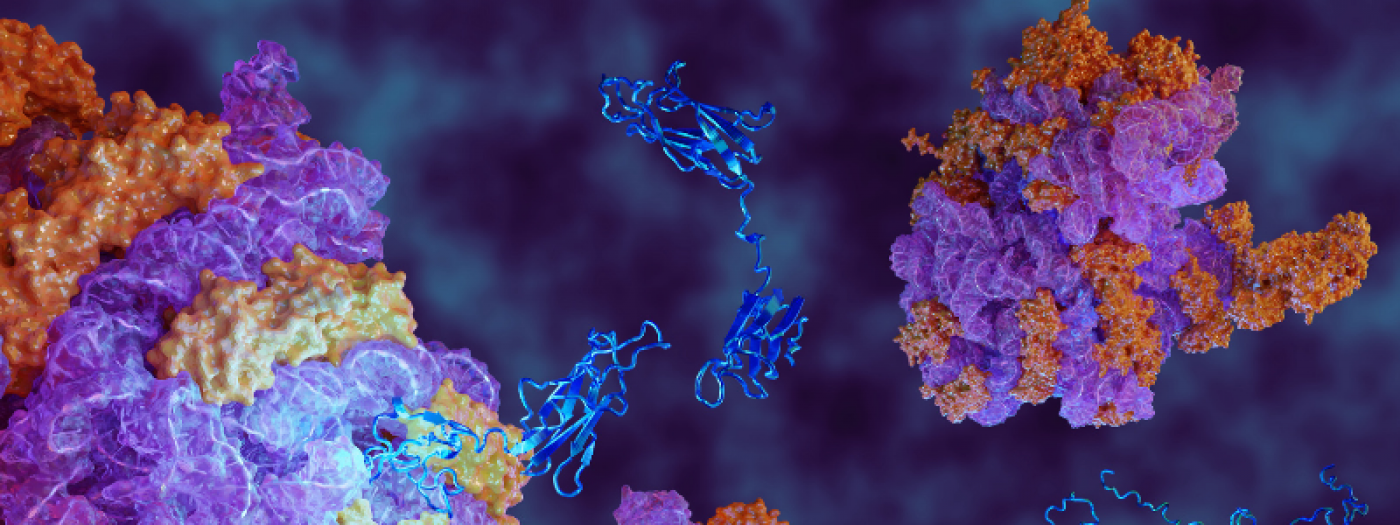
Protein folding in health and disease
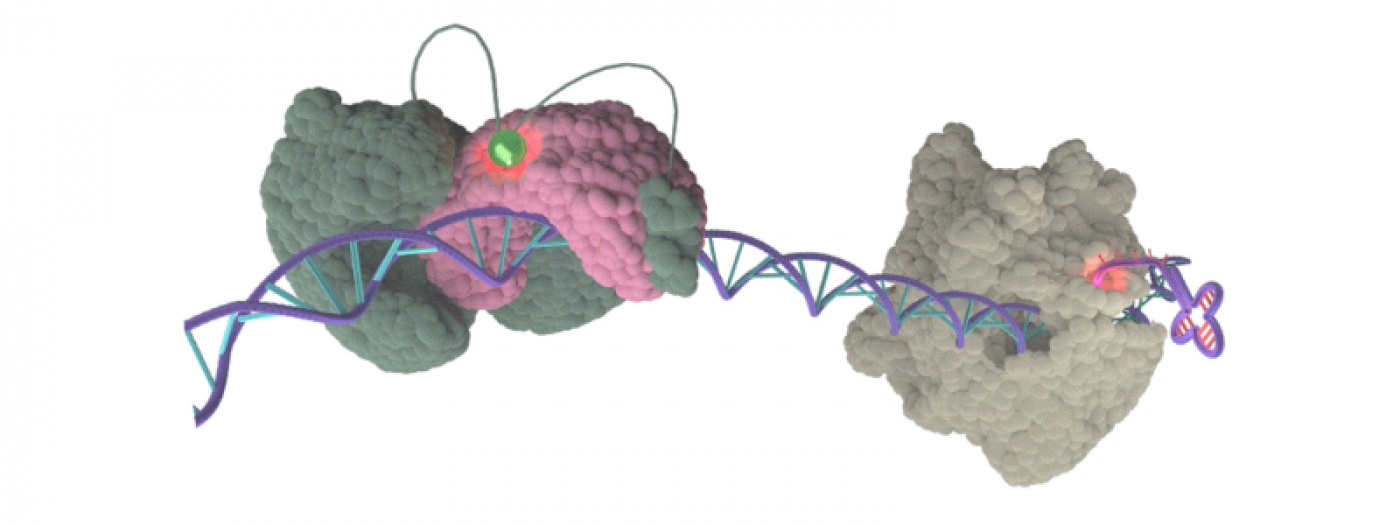
Transcription and RNA Biology
Study


Find resources and support for promoting student and staff wellbeing

Find out about sustainability initiatives for labs and individuals
Contact Us
Department of Structural and Molecular Biology
Division of Biosciences
UCL Darwin Building
Gower Street
London WC1E 6BT
General Enquiries tel: +44 (0)20 7679 2308, email: tabitha.owen@ucl.ac.uk
Postgraduate Enquiries email: biosciences.pgr.admin@ucl.ac.uk
Undergraduate Enquiries email: biosciences-admissions@ucl.ac.uk
The majority of our laboratories and research spaces are located in the UCL Darwin Building on Gower Street, formerly the site of 'Macaw Cottage', a residence of Charles Darwin's.
The main reception to the building is accessed via Darwin Walk, off Malet Place, entered via Torrington Road.
UCL Map for Darwin Building entrance
 Close
Close