Group Leader: Professor Marc R Mansour, MBChB PhD MRCP FRCPath
GOS Children's Charity Professor of Haematology, UCL Cancer Institue and UCL Great Ormond Street Institute of Child Health
Honorary Consultant Haematologist, UCLH.
Research overview
Our laboratory focuses on the biology of two highly aggressive haematological malignancies, namely T-cell acute lymphoblastic leukaemia (T-ALL) and acute myeloid leukaemia (AML).
1) Oncogenic enhancers: we are interested in characterising the mechanisms that lead to malignant transformation in T-ALL and AML, particularly in how somatic mutations in non-coding (or ‘junk’) DNA can activate oncogenes. These gain-of-function non-coding mutations, that we term ‘NEMOs’ (Neomorphic Enhancer Mutations of Oncogenes), create new binding sites for transcription factors, recruit histone modifying machinery and form enhancers to activate aberrant expression of neighbouring oncogenes. Through genome editing, we have been able to show that non-coding mutations, such as those driving TAL1 expression in T-ALL, are absolutely required for oncogene expression and consequently leukaemia cell survival (Figure 1). Mechanisms of aberrant enhancer formation in malignancy have broad implications not only for molecular pathogenesis, but also for clinical management. Drugs that target key components of the transcriptional machinery, such as BRD4 and CDK7 (Figure 1), have recently been shown to preferentially target tumor-specific enhancers, providing a novel strategy to capitalise on these abnormalities for improved cancer therapy
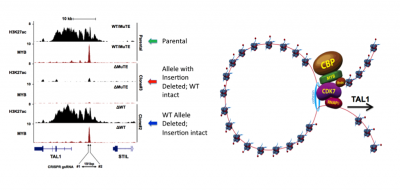
Figure 1. Somatic mutations in non-coding DNA drive aberrant enhancer formation and are potentially targetable with drugs.
Shown are ChIP-seq tracks for MYB and the activation mark H3K27ac in Jurkat T-ALL cells, which harbour a 12 bp insertion in a non-coding site that introduces a new binding motif for MYB (Mansour et al. Science 2014). CRISPR/Cas9-mediated deletion of the wild-type allele has no effect on the enhancer, while deletion of the mutant allele leads to complete collapse of the oncogenic enhancer. The non-coding mutations at oncogenes such as TAL1 lead to aberrant enhancer loop formation and recruitment of abundant CDK7 and BRD4, which are now specifically targetable with drugs.
2) Synthetic lethality: traditional approaches to the development of new treatments for cancer have focused on targeting oncogenes. Such therapies are often limited by resistance mechanisms and dose-limiting effects on normal cells. Given most cancers are also characterised by loss of tumour suppressor genes, either through genetic deletion, mutation or methylation, one potential approach to specific cancer therapy is to exploit the concept of synthetic lethality (Figure 2). This idea was first developed in the study of Drosophila by Calvin Bridges in 1922, but is likely to be highly relevant to modern cancer therapeutics:
‘Two genes are synthetic lethal if mutation of either alone is compatible with viability but mutation of both leads to death. Thus, targeting a gene that is synthetic lethal to a cancer-relevant mutation should kill only cancer cells and spare normal cells.’ Professor William Kaelin, Nature Reviews Cancer.
Cancer cell lines have developed many thousands of mutations during their evolution in culture, making them a suboptimal model for studying synthetic lethality. Instead, we plan to harness the power of the zebrafish to study cancer relevant gene mutations in vivo. Using CRISPR/Cas9 genome engineering, we are creating zebrafish lines harbouring loss-of-function mutations in genes known to be recurrently deleted/mutated in leukaemia. Given a single zebrafish pair is capable of producing several hundred eggs per week, it should be possible to test many thousands of drugs in a relatively short period of time, for their ability to kill the pre-malignant cells of mutant fish, while leaving normal fish embryos unaffected. Thus we hope to identify drugs that are synthetic lethal with cancers cells lacking a specific tumour suppressor, while leaving normal cells unharmed.
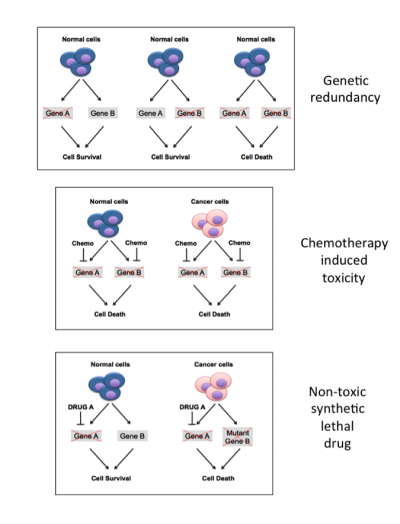
Figure 2. Principles of synthetic lethality
Two genes, A and B, are said to be redundant if cells can survive in the absence of one, or the other, but not both. Standard chemotherapy agents have widespread effects on cells, and are toxic to both cancer cells and normal cells. Most cancers harbour one or more mutation that inactivates a tumour suppressor gene, shown in the figure here as gene B. Thus identifying drugs that can specifically target gene A offers the opportunity to cause death of cancer cells, while leaving normal cells unharmed.
Selected publications
O'Connor D, Espejo Valle-Inclan J, Conde L, Bloye G, Rahman S, Costa JR, Bartram J, Adams SP, Wright G, Elrick H, Wall K, Dyer SA, Howell CD, Jigoulina G, Herrero J, Cortes-Ciriano I, Moorman AV, Mansour MR: Non-coding mutations drive persistence of a founder pre-leukemic clone which initiates late relapse in T-ALL. Blood 2024.
O'Connor D, Demeulemeester J, Conde L, Kirkwood A, Fung K, Papaleonidopoulou F, Bloye G, Farah N, Rahman S, Hancock J, Bateman C, Inglott S, Mee J, Herrero J, Van Loo P, Moorman AV, Vora A, Mansour MR: The Clinicogenomic Landscape of Induction Failure in Childhood and Young Adult T-Cell Acute Lymphoblastic Leukemia. J Clin Oncol 2023, 41:3545-56.
O'Brien H, Salm M, Morton L, Szukszto M, O'Farrell F, Boulton C, Becker PD, Samuels Y, Swanton C, Mansour MR, Reker Hadrup S, Quezada SA: Breaking the performance ceiling for neoantigen immunogenicity prediction. Nat Cancer 2023, 4:1618-21.
Maciocia, P. M., Wawrzyniecka, P. A., Maciocia, N. C., Burley, A., Karpanasamy, T., Devereaux, S., Hoekx, M., O'Connor, D., Leon, T. E., Rapoz-D'Silva, T., Pocock, R., Rahman, S., Gritti, G., Yanez, D. C., Ross, S., Crompton, T., Williams, O., Lee, L. S. H., Pule, M. & Mansour, M. R. Anti-CCR9 Chimeric Antigen Receptor T cells for T Cell Acute Lymphoblastic Leukemia. Blood, doi:10.1182/blood.2021013648 (2022).
Llombart, V. & Mansour, M. R. Therapeutic targeting of "undruggable" MYC. EBioMedicine 75, 103756, doi:10.1016/j.ebiom.2021.103756 (2022).
 Close
Close


